Proteomic characterization of the world trade center dust-activated mdig and c-myc signaling circuit linked to multiple myeloma
- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:
ABSTRACT Several epidemiological studies suggested an increased incidence rate of multiple myeloma (MM) among first responders and other individuals who exposed to World Trade Center (WTC)
dust. In this report, we provided evidence showing that WTC dust is potent in inducing mdig protein and/or mRNA in bronchial epithelial cells, B cells and MM cell lines. An increased mdig
expression in MM bone marrow was observed, which is associated with the disease progression and prognosis of the MM patients. Through integrative genomics and proteomics approaches, we
further demonstrated that mdig directly interacts with c-myc and JAK1 in MM cell lines, which contributes to hyperactivation of the IL-6-JAK-STAT3 signaling important for the pathogenesis of
MM. Genetic silencing of mdig reduced activity of the major downstream effectors in the IL-6-JAK-STAT3 pathway. Taken together, these data suggest that WTC dust may be one of the key
etiological factors for those who had been exposed for the development of MM by activating mdig and c-myc signaling circuit linked to the IL-6-JAK-STAT3 pathway essential for the
tumorigenesis of the malignant plasma cells. SIMILAR CONTENT BEING VIEWED BY OTHERS COAL DUST NANOPARTICLES INDUCED PULMONARY FIBROSIS BY PROMOTING INFLAMMATION AND EPITHELIAL-MESENCHYMAL
TRANSITION VIA THE NF-ΚB/NLRP3 PATHWAY DRIVEN BY IGF1/ROS-MEDIATED AKT/GSK3Β SIGNALS Article Open access 29 December 2022 HIGH EXPRESSION OF ITGB3 AMELIORATES ASTHMA BY INHIBITING
EPITHELIAL-MESENCHYMAL TRANSFORMATION THROUGH SUPPRESSING THE ACTIVATION OF NF-KB PATHWAY Article Open access 22 April 2025 ASSESSMENT OF TOXICITY CHANGES INDUCED BY EXPOSURE OF HUMAN CELLS
TO LUNAR DUST SIMULANT Article Open access 21 October 2024 INTRODUCTION Multiple myeloma (MM) is a malignant neoplasm of plasma cells localized within the bone marrow (BM) compartment and
ranked second in prevalence of all hematopoietic malignancies1. In 2014, there were around 24,000 and 110,000 new cases in U.S and worldwide, respectively. Despite extensive effort was made
in the past decades, the risk factors for MM are still elusive. Several earlier studies suggested that environmental exposures to industrial or agricultural products, such as benzene,
petroleum products, and pesticides, may contribute to the development of MM2,3. Some recent cohort studies on the first responders, reconstruction workers and volunteers of the World Trade
Center (WTC) after terrorist attack on September 11, 2001, provided evidence linking inhalation of the WTC dust to MM and other types of malignancy4,5,6. However, there are no studies
revealing whether WTC dust is potentially carcinogenic, and if so, how WTC dust causes malignant transformation of the mature plasma B cells. In the past decade, large-scale genomics studies
have determined genetic landscape of MM and identified abnormal genetic events present in various disease stages, from premalignant monoclonal gammopathy of undetermined significance (MGUS)
to smoldering multiple myeloma (SMM), active MM and relapsed MM7. The unique BM milieu is vital for the longevity of myeloma cells by providing various supportive BM cells and soluble
factors1. Among these external stimuli, one of the most important factors is the interleukin-6 (IL-6). After binding to its receptor (IL-6R) and recruiting signal transducer, GP130 (also
known as CD130 or IL-6ST), IL-6 can activate Janus Kinase (JAK)/signal transducer and activator of transcription 3 (STAT3), Akt and mitogen-activated protein kinase (MAPK) pathways to
promote proliferation, survival and drug resistance of the MM cells8. Another hallmark of MM pathogenesis is the mutation- or overexpression-induced c-myc activation9. C-myc is a
well-defined onco-protein involved in many types of human cancers. As an essential transcription factor, c-myc upregulates transcription of genes responsible for cell growth, proliferation
and maintenance of cancer cell stemness10. In MM, c-myc overexpression can distinguish active MM from premalignant MGUS. In addition, activated c-myc has been shown to sustain the survival
of myeloma cells11. More interestingly, a recent study indicates that collaboration between IL-6 pathway and c-myc results in significant acceleration of MM pathogenesis12. However, the
underlying mechanisms of this oncogenic interaction remain unclear. As a c-myc-induced protein, mdig (mineral dust-induced gene, also known as mina53, MINA, or NO52) functions as a histidyl
hydroxylase and potentially a lysine-specific demethylase, which regulates gene transcription through modifying tri-methylated lysine 9 residue on histone 3 (H3K9me3)13,14. Consistent with
this function, mdig is found to be exclusively localized in nucleus of various cell types15,16. Some studies have demonstrated that mdig exerts strong immune-regulatory function on T cells
by favoring Th1 and Th17 cell differentiation in Th1/Th2, Th17/Treg balances, respectively17,18,19,20. Overexpression of mdig has been observed in many types of human cancer, including lung
cancer, colon cancer, gastric carcinoma, etc.21,22,23,24,25,26. Meanwhile, mdig has been shown to be able to promote cancer cell proliferation16,21,27. Furthermore, mdig overexpression has
been exclusively observed in various B cell-derived malignancies among major human lymphoma subtypes28, suggesting that mdig may contribute to c-myc-induced tumorigenesis in MM. The WTC dust
from the collapse of World Trade Center building due to terrorist attack is a mixture of toxic materials, many of which are know human carcinogens, including crystalline silica, asbestos,
carbon nanotubes, lead, cadmium, mercury, dioxins, polycyclic aromatic hydrocarbons, etc29,30. By studying cancer incidence rate among workers of rescue, recovery, reconstruction, and
volunteers who were exposed to the heavy dust cloud at Ground Zero and registered in the New York/New Jersey consortium of the WTC Medical Monitoring and Treatment Program (MMTP), Moline _et
al_. noted 8 cases of MM, four of them were younger than 45 years4. This observation was supported by another longitudinal study of 55,778 New York State residents, which revealed excess
incidence in responders for MM5. In the present report, we provide evidence revealing that WTC dust is potent in inducing mdig in normal B cells and MM cells and further demonstrating that
mdig is significantly associated with the malignant transformation of MGUS to active MM, disease exacerbation and poor clinical outcomes. Biochemical studies unraveled that mdig directly
interacts with c-myc and JAK1 in MM cells, which attributes to the hyperactivation of the JAK1 and STAT3 signaling important for cell survival, proliferation and development of drug
resistance of the MM cells31,32. Taken together, our data suggest that mdig may serve as a key mediator for MM associated with WTC dust exposure and potential diagnosis/prognosis marker of
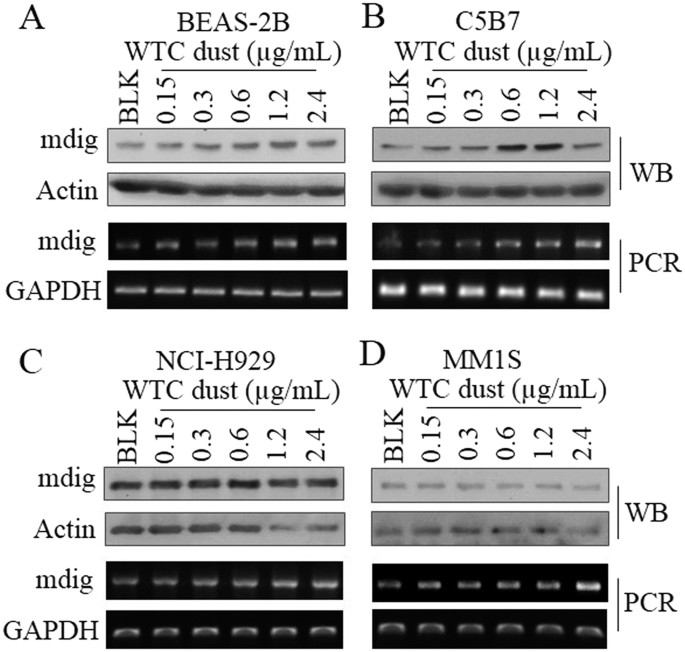
MM. RESULTS WTC DUST INDUCES MDIG IN BRONCHIAL EPITHELIAL CELLS, B CELLS AND MM CELLS The adverse effect of WTC dust on the respiratory system, including airway inflammation, impairment of
the pulmonary function, airway hyperactivity, asthma, and sarcoid-like granulomatous pulmonary disease, had been well-established33,34. Indeed, we noted that WTC dust is highly capable of
inducing mdig expression in the bronchial epithelial cell line, BEAS-2B cells, in concentrations ranged from 0.15 to 2.4 μg/ml (Fig. 1A). Since concerns had been arisen about the potential
for increased risk of MM among WTC responders4,5, we also investigated the capability of WTC dust on the induction of mdig in normal B cells using a B cell line C5B7. Similar to what we
observed in BEAS-2B cells, we noted a dose-dependent induction of mdig protein and mRNA by WTC dust in C5B7 cells (Fig. 1B). In two MM cell lines NCI-H929 and MM1S, although we did not
detect significant induction of mdig protein, a pronounced induction of mdig mRNA by WTC dust was observed (Fig. 1C,D). These data, thus, clearly suggest that in addition to damaging the
respiratory system through direct interaction, WTC dust or its components may influence the function or growth status of the B cells and the MM cells. INCREASED MDIG EXPRESSION IN THE BONE
MARROW (BM) OF THE MM PATIENT To determine whether mdig expression is clinically relevant for MM, we evaluated mdig protein levels in the BM specimens of MM patients through
immunohistochemistry (IHC). In total of 16 cases of MM BM biopsies examined, 8 samples exhibited strong staining of mdig proteins as judged by the fact that more than 50% of cells are mdig
positive, 6 samples showed moderate or weak mdig staining and 2 samples are mdig negative (Fig. 2A,B). We also checked another set of BM specimens collected from 4 healthy donors and 11
patients with other non-hematological cancers. Mdig protein was not detected in the BM specimens of all 4 healthy donors and 8 out of 11 cases of non-hematological cancer patients. Three BM
specimens from patients with non-hematological cancers showed weak positive of mdig staining (Fig. 2A,B). Thus, these data indicate that the level of mdig may be associated with the onset of
MM. BOTH MDIG AND C-MYC ARE ASSOCIATED WITH DISEASE AGGRESSIVENESS OF MM PATIENTS There are several stages during disease development, including premalignant MGUS, asymptomatic smoldering
MM (SMM), symptomatic MM, and relapsed MM35. It has been well-accepted that c-myc activation is a hallmark of MM pathogenesis11,36,37, especially in the early malignant transformation from
MGUS to active MM38. C-myc has also been implicated in the up-regulation of mdig27. Overexpression of mdig has been observed in many types of human malignancies13,14,21,22,23,24,25,26,27,28,
but its potential role in c-myc-related MM pathogenesis remains unknown. To determine whether mdig contributes to c-myc-induced MM pathogenesis, we first examined expression levels of mdig
and c-myc in MM patients. We noted that both c-myc and mdig mRNAs are significantly up-regulated in newly diagnosed MM patients when compared to healthy donors (Fig. 3A,B). Further analysis
of patients at continuous stages during MM development has demonstrated a robust elevation trend of both c-myc and mdig (Fig. 3C,D). Statistically significant increases of mdig mRNA, from
MGUS to active MM and from SMM to relapsed MM were noted (Fig. 3D), suggesting a positive correlation between mdig expression and malignant transformation, disease progression and relapse of
MM. The involvement of mdig in MM pathogenesis is further supported by survival analysis of 559 MM patients39. High level of mdig expression is significantly correlated with poor overall
survival of the MM patients, even though higher percentage of patients from mdigHigh group (82%, 96/117) received intensive therapies than those from mdigLow group (57.7%, 255/442) (Fig.
3E). Taken together, all above data demonstrate a strongly positive correlation of mdig and c-myc to the pathogenesis and aggressiveness of MM. MDIG ACTS AS A KEY INTERACTION PARTNER OF
C-MYC IN MM CELLS In order to decipher inter-regulation between mdig and c-myc in MM cells, proteomics study was performed on MM cell line NCI-H929 cells to screen their interaction
partners, respectively. A total of 224 and 203 proteins were identified as significant binding partners of mdig and c-myc, respectively. Among them, 110 binding partners are shared by mdig
and c-myc (Fig. 4A). Strikingly, physical binding between mdig and c-myc was detected by mass-spectrometry in NCI-H929 cells (Fig. 4B), which was further validated by co-IP assay in both
NCI-H929 and MM1S cells (Fig. 4C), implying that mdig might be assembled into functional protein complexes together with c-myc and directly participate in c-myc-induced oncogenesis for the
development of MM. Subsequent network analysis highlighted some major cellular events upon which c-myc and mdig are most likely to impose their impact (Fig. 4D). The shared binding partners
are mainly clustered in 4 areas: gene expression, post-transcriptional regulation of gene expression, mRNA processing, and mRNA transport. It is not surprising that c-myc-only binding
partners are actively involved in all 4 biological processes and mdig-only binding partners involved in former 2 processes considering the well-established role of c-myc as an essential
transcription factor10,37,40 and mdig as an important epigenetic regulator13,14. Collectively, these data provide a strong rationale that mdig is a core direct interaction partner of c-myc
and is most likely to collaborate in gene expression-related functions in MM cells. Notably, mdig-only binding partners are also enriched in proteins important for cellular responses to
cytokine and antigen processing and presentation, which is in agreement with our previous findings suggesting that mdig contributes to the function of the T helper 17 (Th17) cells17,41. Most
recently, we discovered that mdig interacts with DNA double strand break repair proteins in the non-homologous end-joining (NHEJ) pathway in human bronchial epithelial cells and lung cancer
cells42. In MM cells, we also identified at least 7 DNA repair proteins that interact with mdig, including XRCC5, XRCC6, Rad50, etc. (Fig. 4D), indicating that mdig may also be involved in
handling cellular stress caused by ongoing DNA damage, a common feature in human MM36,43. MDIG BINDS JAK1 IN MM CELLS Among the most important signaling pathways, IL-6/JAK/STAT3 signaling
has been viewed as an indispensable signal for the malignant transformation of plasma B cells and proliferation of the MM cells44. Through cooperation with c-myc, this signaling pathway
drives formation of highly malignant MM in mouse model45. It is unclear how this cooperation is established between the oncogenic signal and cytokine signal. It is noteworthy that proteomic
study identifies JAK1, a key regulator mediating cytokine-induced signaling, as a significant interaction partner of mdig (Fig. 5A). Following this clue, co-IP assay was performed using
total cell lysates of both NCI-H929 and MM1S cells and confirmed such a physical interaction (Fig. 5B). The interaction of mdig and JAK1 was additionally verified by immunofluorescent
staining and confocal microscopy. Multiple co-localization sites of mdig and JAK1 were observed in the extra-nuclear area in both NCI-H929 cells and MM1S cells (Fig. 5C). Mdig has long been
recognized as a nuclear protein15,16, whereas JAK1 is believed to be a cytosolic protein in the proximity of cytokine receptors. It is interesting to know how a nuclear protein can interact
with a cytosolic protein. To answer this question, different cellular compartments were separated through fractionation. Surprisingly, in both MM cell lines, a significant portion of mdig
was found in cytosol though the majority of mdig located in nucleus (Fig. 5D). Thus, cytosolic localization of mdig may be accounted for the proximity and physical interaction between mdig
and JAK1. This is also the first observation of mdig in cytosol of human cell lines without additional manipulation, although we had also noted cytosolic localization of mdig in
mdig-overexpressed or arsenic-treated A549 cells16. MDIG STABILIZES JAK1 To investigate the biological function of mdig-JAK1 interaction, we further studied the role of mdig on the gene
expression and protein stability of the JAK1 protein in MM cells. The co-amplification analysis on MM patients exhibits no significant difference of JAK1 mRNA level between “mdig high” and
“mdig low” groups (Fig. 6A). In NCI-H929 cells, genetic silencing of mdig does not affect mRNA level of JAK1 (Fig. 6B), while in MM1S cells, mdig knock-down groups displayed slightly higher
JAK1 mRNA expression than the control group (Fig. 6C). However, on the protein level, silencing mdig resulted in a considerable decrease of total JAK1 protein (Fig. 6D). These data suggest
that mdig affects the JAK1 protein level through some posttranslational mechanisms. Given the potential activity of mdig on lysine demethylation13,14, we hypothesize that mdig may regulate
JAK1’s stability by removing the methyl groups from its lysine residue(s). Because there is no report of JAK1 methylation so far and the unavailability of antibodies targeting methylated
JAK1, we first immunoprecipitated and collected JAK1 protein from the control and mdig-silenced MM cells and then probed the samples with an antibody that selectively recognizes methylated
lysine. We showed a marginal lysine methylation on JAK1 in both NCI-H929 and MM1S cells in which mdig was silenced by siRNA (data not shown). However, this set of data remains to be further
confirmed. MDIG AND C-MYC ARE REQUIRED FOR THE HYPERACTIVATION OF THE IL-6 SIGNALING Synergetic collaborations between c-myc and IL-6 pathways have been well-documented in MM12,37,46.
Prompted by the implications from proteomics studies above, we next interrogated the possibility of mdig in mediating the oncogenic crosstalk between c-myc and IL-6 signaling. Consistent
with a previous report47, our biochemical analysis demonstrated that genetic silencing of mdig results in decreased protein levels of GP130, but not IL-6R in both NCI-H929 and MM1S cell
lines (Fig. 7A). Moreover, mdig silencing further leads to attenuated phosphorylation of the major downstream effectors on IL-6 signaling pathway, including STAT3 on both Tyrosine 705 and
Serine 727 sites, and Akt on Serine 473 site, but not their total protein levels (Fig. 7A). On the other hand, silencing c-myc or treatment of the cells with a c-myc inhibitor, 10058-F4,
leads to a notable decrease of the mdig protein and all regulators on IL-6 pathway mentioned above, except pAkt (Fig. 7B,C), indicating that c-myc is an essential transcription factor in MM
cells while mdig specifically cooperates with c-myc in promoting overexpression of GP130 and, consequently, causes amplification of the IL-6 signaling for cell survival and growth.
DISCUSSION Considerable progresses in understanding the molecular pathogenesis of MM have been achieved in the past years. However, many important questions remain to be answered, such as
the risk factors for MM and the extensive crosstalk between various oncogenic mechanisms in MM. Bone marrow is a complex and dynamic microenvironment with stromal cells, osteoclasts, T
lymphocytes, cytokines and growth factors, which are critical for disease evolution of MM. In such a profoundly-intertwined regulatory network of malignancy, oncogene c-myc and cytokine IL-6
have long been viewed as major driving forces for the pathogenesis of MM36,48,49. In the present study, we provide the first evidence showing that WTC dust is potent in inducing mdig in
normal B cells and MM cells, and mdig is a key mediator in synergizing c-myc and IL-6 signaling through direct interaction with c-myc and JAK1. By both upregulating and sustaining key
regulators in IL-6 pathway, mdig enables MM cells to take advantage of this critical intracellular pathway to achieve abnormal cell proliferation and apoptosis escape. These results explain,
at least in part, the mechanisms underlying the observed synergetic collaboration between IL-6 pathway and c-myc in promoting oncogenesis of the plasma cells. A study by Moline _et al_.4
suggested an increased incidence rate and early onset of MM among the first responders who exposed to the WTC dust. A follow-up study by Li _et al_.5 on 55,778 people, including rescue
workers, recovery workers, and those who lived or worked near the WTC, also found a higher rate of MM, in addition to thyroid and prostate cancers. Both of these studies noted that there
were no previous reports indicating association of special occupation, such as those firefighters and police officers, with MM. The WTC dust released from the collapse of the twin towers
after 9/11 attack is a mixture of mineral particles, fibers, metals, and chemicals, many of which are established human carcinogens30. Since mdig was originally identified as a mineral
dust-induced gene from coal workers who exposed to mining and coal dust in a daily basis21, we sought to determine whether induction of mdig can be indicative for the association of MM and
WTC dust. Indeed, we found that WTC dust is highly capable of inducing mdig expression in either bronchial epithelial cells, normal B cells, or the MM cells. Although the results reported
here can be viewed as circumstantial, they may be considered as “proof of principle” to address the carcinogenic potential of environmental factors on the development of MM. The findings
that mdig is strongly associated with the disease progression of MM patients suggest that mdig can be potentially used as a prognostic marker to guide clinical management of the MM patients.
A similar role of mdig had been reported in human gastric carcinoma50. Our analysis shows that mdig mRNA significantly increases as disease progresses. Notably, the increases of mdig
expressions in MM verse MGUS and MM versus SMM are both statistically significant. In addition, high level of mdig is also significantly associated with poor overall survival of the MM
patients. In the cellular models, we have demonstrated that genetic silencing of mdig in MM cells leads to constitutive suppression of GP130 (IL-6ST) and pro-survival regulators, STAT3 and
Akt, suggesting that mdig inhibition could be a possible strategy to suppress tumor growth in IL-6-dependent MM subtypes or sensitize them to IL-6-targeted agents. The mdig protein contains
a conserved JmjC domain without classic chromatin- or DNA-binding domains14,51. In accordance with a recent report42, our proteomic analysis has unraveled direct interactions of mdig with a
number of chromatin-binding proteins and DNA repair proteins. Given that our current findings have clearly demonstrated a regulatory circuit among c-myc, mdig and IL-6 signaling, it is
plausible to speculate that mdig may be assembled into protein complexes with chromatin- or DNA-binding protein(s), like c-myc, and be recruited to MM-specific signature genes, including
GP130, and exerts its regulatory functions on gene expression. On the other hand, recent studies have discovered that transcription-related regulators can translocate to different cellular
compartments and carry out non-canonical functions. For example, Enhancer of Zeste Homolog 2 (EZH2), a well-documented epigenetic silencer for gene transcription, has been shown to directly
interact with and methylate STAT352. Similarly, in the present report, we have observed that mdig binds to JAK1 in cytosol, which may stabilizes JAK1 protein. In summary, we provided the
first evidence showing that WTC dust is highly capable of inducing mdig expression in human bronchial epithelial cells, normal human B cells and human MM cell lines. The levels of mdig mRNA
and protein are associated with the disease progression and prognosis of the MM patients. Proteomic analyses suggested a direct interaction of mdig with c-myc and JAK1 in MM cell lines,
which was responsible for the hyperactivation of the IL-6-JAK-STAT3 signaling important for the pathogenesis of MM. Additional studies are much needed to determine whether the level of mdig
expression can be used to guide the clinical managements of the MM. MATERIALS AND METHODS CELLS AND REAGENTS Human MM cell lines, NCI-H929 and MM1S, bronchial epithelial cell line BEAS-2B,
and normal B cell line C5B7 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and maintained in ATCC-recommended culture conditions. Inhibitor of c-myc
(10058-F4) was purchased from Sigma-Aldrich Co. (St. Louis, MO, USA). WTC dust was provided by Dr. Kenneth Reuhl at the Environmental and Occupational Health Sciences Institute of the
Rutgers University. SIRNA TRANSFECTION Transfections were performed using Lipofectamine RNAiMAX (Invitrogen) according to manufacturer’s protocol. Fifty nM of siRNAs were used for
transfection followed by 48-hour incubation. Control siRNA, Mdig siRNAs and c-myc siRNAs were purchased from Qiagen (Valencia, CA, USA). IMMUNOHISTOCHEMISTRY (IHC) Tissue microarray slides,
T293 and BM483b, containing multiple myeloma samples and non-cancerous bone marrow tissue were purchased from US Biomax, Inc (Rockville, MD). IHC staining was performed as previously
described23. Briefly, the slides were stained overnight at 4 °C with mouse anti-human mdig antibody (Invitrogen) at 1:50 dilution followed by biotinylated goat anti-mouse secondary antibody
(Dako Denmark A/S, Glostrup, Denmark) at 1:200 dilution for 2 hours at room temperature. The slides were then incubated with ABC reagent and DAB (Vector Laboratories, Inc. Burlingame, CA),
counter stained with hematoxylin and mounted with entellan. All images were captured using a Nikon Eclipse Ti-S Inverted microscope (Mager Scientific, Dexter, MI). Cut-offs between positive
and negative cells was determined according to previously characterized mdig-expressing breast cancer samples23. Four random images were taken for each sample and both positive and negative
cells were counted using ImageJ 1.48 v ( http://imagej.nih.gov/ij/). Mdig expression status of all samples was classified into four grades based on the percentage of positively-stained
cells. Strongly positive: over 50%; moderately positive: between 50% and 25%; weakly positive: between 25% and 5%; negative: less than 5%. IMMUNOBLOTTING AND IMMUNOPRECIPITATION (IP)
Immunoblotting and IP analysis were performed as previously reported42. NE-PER Nuclear Cytoplasmic Extraction KIT (Thermo Scientific Pierce, Rockford, IL, USA) was used to isolate nuclear
proteins. Densitometric analysis of CHX-treated samples was completed using ImageJ 1.48 v ( http://imagej.nih.gov/ij/). When detecting c-myc bands in IP samples, HRP-conjugated protein A
(EMD Millipore, Temecula, CA, USA) was used to minimize the background noise caused by IgG heavy chain. Primary antibodies against phospho-Akt (Ser473), total Akt, phospho-STAT3 (Ser727),
phospho-STAT3 (Tyr705), total STAT3, phospho-JAK1 (Tyr1022), total JAK1, GAPDH, actin and all secondary antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA).
Antibodies against GP130, IL-6R and methylated-lysine were purchased from Abcam (Cambridge, MA). Antibodies against c-myc and lamin A/C were purchased from Santa Cruz Biotechnology (Dallas,
Texas, USA). Mdig (mouse) antibody was ordered from Invitrogen. Antibodies used for IP include Mdig (rabbit) and c-myc (mouse) from Abcam (Cambridge, MA, USA), c-myc (rabbit) from Cell
Signaling Technology (Danvers, MA, USA), JAK1 (rabbit) from Santa Cruz Biotechnology (Dallas, Texas, USA). All presented data are representatives of at least 3 independent experiments.
CONFOCAL IMMUNOFLUORESCENCE (IF) ANALYSIS For IF staining, 106 cells were centrifuged, fixed by 4% formaldehyde for 15 min, permeabilized by 0.3% Triton X-100 and blocked in PBS containing
5% normal goat serum and 0.1% Tween 20 for 1 hour at room temperature. Then they were incubated with primary antibodies, JAK1 (rabbit, Santa Cruz Biotechnology) and Mdig (mouse, Invitrogen)
overnight at 4 °C and with Invitrogen secondary antibodies, Alexa Fluor 488-linked antibody (goat anti-mouse) and Alexa Fluor 594-linked antibody (goat anti-rabbit) for 1 h at room
temperature in dark. All antibodies were used at 1:100 dilution. Prolong Gold antifade reagent with DAPI (Invitrogen) was used to preserve the samples. Co-localization of JAK1 and Mdig was
detected by Zeiss LSM 780 confocal microscope (Carl Zeiss Microscopy, Jena, Germany). Pinhole size of 60 μm was used while thresholds for laser power, master gain and digital gain were
determined by non-specific binding controls. DAPI, Alexa Fluor 488 and Alexa Fluor 594 were excited at 405 nm, 488 nm and 595 nm and corresponding fluorescence emissions were detected at 495
nm, 563 nm and 640 nm via 3 independent channels. All photos were processed using ZEN 2012 SP1 64 bit software (Carl Zeiss Microscopy, Jena, Germany). PCR Total RNAs were extracted using
TRIzol Reagent (Life Technologies, Grand Island, NY, USA) and their integrity was assessed by 18S and 28S ribosomal RNAs. For reverse transcription PCR, AccessQuick RT-PCR system from
Promega (Madison, WI) was used. The primers for mdig are: 5′-TCA TGT CGG GCC TAA GAG AC-3′ and 5′-GGC ATT TGA TTC TGC AAA GG-3′, which amplifies a 1,510 bp DNA fragment covering the whole
coding region of the mdig gene. Primers for GAPDH are: 5′-CTG AAC GGG AAG CTC ACT GGC ATG GCC TTC-3′ and 5′-CAT GAG GTC CAC CAC CCT GTT GCT GTA GCC-3′. For real-time PCR, one μg total RNAs
were reverse-transcribed using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Waltham, MA, USA) and 1:20 diluted. Jak1 and ACTB Taqman Gene Expression Assays (Best
Coverage) were purchased from Applied Biosystems (Waltham, MA, USA). Samples were run in triplicates, quantified by ΔΔCt method with actin as reference gene and normalized to “Blank” group.
Final results were shown as mean ± SD. MASS SPECTROMETRY AND PROTEOMICS ANALYSIS Proteomics profiling of binding partners were performed as previously reported42. Briefly, samples were
subject to co-IP, 1D-SDS-PAGE separation, in-gel digestion, peptide purification and HPLC-ESI-MS/MS analysis. Protein identity was determined by MaxQuant software. BIOSTATISTICS ANALYSIS
Protein interaction network analysis was completed using Gene Ontology database and visualized by Cytoscape 3.2. Binding proteins were first sorted according to their biological processes
and further refined manually by merging repeating and redundant categories. Gene expression data were accessed through Multiple Myeloma Genomics Portal (
https://www.broadinstitute.org/mmgp/home) for GSE6477 and through GEO for GSE39754 and GSE2658 before being processed and visualized using R project with ggplot2 package. Survival analysis
in Fig. 3E was performed using Kaplan-Meier method and differences between 2 cohorts were determined using log-rank test. In Fig. 3C,D, differences of mRNA levels between patient cohorts
were calculated using one-way ANOVA and p-values were adjusted by Holm method. All other mRNA expression comparisons were performed using two-tailed t-test. Considering that expression
levels of related genes are not always strictly linear to each other, we conducted “Force Rank” co-amplification analysis as previously described53. A p-value less than 0.05 is considered
statistically significant. ADDITIONAL INFORMATION HOW TO CITE THIS ARTICLE: Wu, K. _et al_. Proteomic Characterization of the World Trade Center dust-activated mdig and c-myc signaling
circuit linked to multiple myeloma. _Sci. Rep._ 6, 36305; doi: 10.1038/srep36305 (2016). PUBLISHER’S NOTE: Springer Nature remains neutral with regard to jurisdictional claims in published
maps and institutional affiliations. REFERENCES * Ribatti, D., Nico, B. & Vacca, A. Importance of the bone marrow microenvironment in inducing the angiogenic response in multiple
myeloma. Oncogene 25, 4257–4266, 10.1038/sj.onc.1209456 (2006). Article CAS PubMed Google Scholar * Vlaanderen, J., Lan, Q., Kromhout, H., Rothman, N. & Vermeulen, R. Occupational
benzene exposure and the risk of lymphoma subtypes: a meta-analysis of cohort studies incorporating three study quality dimensions. Environmental health perspectives 119, 159–167,
10.1289/ehp.1002318 (2011). Article CAS PubMed Google Scholar * Weichenthal, S., Moase, C. & Chan, P. A review of pesticide exposure and cancer incidence in the Agricultural Health
Study cohort. Environmental health perspectives 118, 1117–1125, 10.1289/ehp.0901731 (2010). Article CAS PubMed PubMed Central Google Scholar * Moline, J. M. et al. Multiple myeloma in
World Trade Center responders: a case series. J Occup Environ Med 51, 896–902, 10.1097/JOM.0b013e3181ad49c8 (2009). Article ADS PubMed Google Scholar * Li, J. et al. Association between
World Trade Center exposure and excess cancer risk. JAMA 308, 2479–2488, 10.1001/jama.2012.110980 (2012). Article CAS PubMed Google Scholar * Solan, S. et al. Cancer incidence in world
trade center rescue and recovery workers, 2001–2008. Environmental health perspectives 121, 699–704, 10.1289/ehp.1205894 (2013). Article PubMed PubMed Central Google Scholar * Shain, K.
H., Dalton, W. S. & Tao, J. The tumor microenvironment shapes hallmarks of mature B-cell malignancies. Oncogene 34, 4673–4682, 10.1038/onc.2014.403 (2015). Article CAS PubMed PubMed
Central Google Scholar * Pene, F. et al. Role of the phosphatidylinositol 3-kinase/Akt and mTOR/P70S6-kinase pathways in the proliferation and apoptosis in multiple myeloma. Oncogene 21,
6587–6597, 10.1038/sj.onc.1205923 (2002). Article CAS PubMed Google Scholar * Cobbold, L. C. et al. Upregulated c-myc expression in multiple myeloma by internal ribosome entry results
from increased interactions with and expression of PTB-1 and YB-1. Oncogene 29, 2884–2891, 10.1038/onc.2010.31 (2010). Article CAS PubMed Google Scholar * Eilers, M. & Eisenman, R.
N. Myc’s broad reach. Genes Dev 22, 2755–2766, 10.1101/gad.1712408 (2008). Article CAS PubMed PubMed Central Google Scholar * Holien, T., Vatsveen, T. K., Hella, H., Waage, A. &
Sundan, A. Addiction to c-MYC in multiple myeloma. Blood 120, 2450–2453, 10.1182/blood-2011-08-371567 (2012). Article CAS PubMed Google Scholar * Rutsch, S. et al. IL-6 and MYC
collaborate in plasma cell tumor formation in mice. Blood 115, 1746–1754, 10.1182/blood-2009-08-237941 (2010). Article CAS PubMed PubMed Central Google Scholar * Chen, B. et al. Mdig
de-represses H19 large intergenic non-coding RNA (lincRNA) by down-regulating H3K9me3 and heterochromatin. Oncotarget 4, 1427–1437 (2013). PubMed PubMed Central Google Scholar * Lu, Y. et
al. Lung cancer-associated JmjC domain protein mdig suppresses formation of tri-methyl lysine 9 of histone H3. Cell Cycle 8, 2101–2109 (2009). Article CAS Google Scholar * Eilbracht, J.,
Kneissel, S., Hofmann, A. & Schmidt-Zachmann, M. S. Protein NO52–a constitutive nucleolar component sharing high sequence homologies to protein NO66. Eur J Cell Biol 84, 279–294,
10.1016/j.ejcb.2004.12.022 (2005). Article CAS PubMed Google Scholar * Yu, M. et al. Paradoxical roles of mineral dust induced gene on cell proliferation and migration/invasion. PLoS One
9, e87998, 10.1371/journal.pone.0087998 (2014). Article ADS CAS PubMed PubMed Central Google Scholar * Thakur, C. et al. Oncoprotein mdig contributes to silica-induced pulmonary
fibrosis by altering balance between Th17 and Treg T cells. Oncotarget 6, 3722–3736 (2015). Article Google Scholar * Bettelli, E. et al. Reciprocal developmental pathways for the
generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238, 10.1038/nature04753 (2006). Article ADS CAS Google Scholar * Yosef, N. et al. Dynamic regulatory
network controlling TH17 cell differentiation. Nature 496, 461–468, 10.1038/nature11981 (2013). Article ADS CAS PubMed PubMed Central Google Scholar * Hemmers, S. & Mowen, K. A.
T(H)2 bias: Mina tips the balance. Nat Immunol 10, 806–808, 10.1038/ni0809-806 (2009). Article CAS PubMed Google Scholar * Zhang, Y. et al. The Human mineral dust-induced gene, mdig, is
a cell growth regulating gene associated with lung cancer. Oncogene 24, 4873–4882, 10.1038/sj.onc.1208668 (2005). Article CAS PubMed Google Scholar * Teye, K. et al. Increased expression
of a Myc target gene Mina53 in human colon cancer. Am J Pathol 164, 205–216, 10.1016/S0002-9440(10)63111-2 (2004). Article CAS PubMed PubMed Central Google Scholar * Thakur, C. et al.
Increased expression of mdig predicts poorer survival of the breast cancer patients. Gene 535, 218–224, 10.1016/j.gene.2013.11.031 (2014). Article CAS PubMed Google Scholar * Tsuneoka,
M. et al. Mina53 as a potential prognostic factor for esophageal squamous cell carcinoma. Clin Cancer Res 10, 7347–7356, 10.1158/1078-0432.CCR-03-0543 (2004). Article CAS PubMed Google
Scholar * Zhang, Q. et al. Expression of Mina53 and its significance in gastric carcinoma. Int J Biol Markers 23, 83–88 (2008). Article CAS Google Scholar * Ishizaki, H. et al.
Overexpression of the myc target gene Mina53 in advanced renal cell carcinoma. Pathol Int 57, 672–680, 10.1111/j.1440-1827.2007.02156.x (2007). Article CAS PubMed Google Scholar *
Tsuneoka, M., Koda, Y., Soejima, M., Teye, K. & Kimura, H. A novel myc target gene, mina53, that is involved in cell proliferation. J Biol Chem 277, 35450–35459, 10.1074/jbc.M204458200
(2002). Article CAS PubMed Google Scholar * Teye, K. et al. Expression of Myc target gene mina53 in subtypes of human lymphoma. Oncol Rep 18, 841–848 (2007). CAS PubMed Google Scholar
* Wu, M. et al. Case report: Lung disease in World Trade Center responders exposed to dust and smoke: carbon nanotubes found in the lungs of World Trade Center patients and dust samples.
Environmental health perspectives 118, 499–504, 10.1289/ehp.0901159 (2010). Article CAS PubMed Google Scholar * Lioy, P. J. et al. Characterization of the dust/smoke aerosol that settled
east of the World Trade Center (WTC) in lower Manhattan after the collapse of the WTC 11 September 2001. Environmental health perspectives 110, 703–714 (2002). Article CAS Google Scholar
* Catlett-Falcone, R. et al. Constitutive activation of Stat3 signaling confers resistance to apoptosis in human U266 myeloma cells. Immunity 10, 105–115 (1999). Article CAS Google
Scholar * Hideshima, T., Nakamura, N., Chauhan, D. & Anderson, K. C. Biologic sequelae of interleukin-6 induced PI3-K/Akt signaling in multiple myeloma. Oncogene 20, 5991–6000,
10.1038/sj.onc.1204833 (2001). Article CAS PubMed Google Scholar * Lippmann, M., Cohen, M. D. & Chen, L. C. Health effects of World Trade Center (WTC) Dust: An unprecedented
disaster’s inadequate risk management. Crit Rev Toxicol 45, 492–530, 10.3109/10408444.2015.1044601 (2015). Article CAS PubMed PubMed Central Google Scholar * Herbert, R. et al. The
World Trade Center disaster and the health of workers: five-year assessment of a unique medical screening program. Environmental health perspectives 114, 1853–1858 (2006). Article Google
Scholar * Korde, N., Kristinsson, S. Y. & Landgren, O. Monoclonal gammopathy of undetermined significance (MGUS) and smoldering multiple myeloma (SMM): novel biological insights and
development of early treatment strategies. Blood 117, 5573–5581, 10.1182/blood-2011-01-270140 (2011). Article CAS PubMed PubMed Central Google Scholar * Bianchi, G. & Munshi, N. C.
Pathogenesis beyond the cancer clone(s) in multiple myeloma. Blood 125, 3049–3058, 10.1182/blood-2014-11-568881 (2015). Article CAS PubMed PubMed Central Google Scholar * Morgan, G. J.,
Walker, B. A. & Davies, F. E. The genetic architecture of multiple myeloma. Nat Rev Cancer 12, 335–348, 10.1038/nrc3257 (2012). Article CAS PubMed Google Scholar * Chng, W. J. et
al. Clinical and biological implications of MYC activation: a common difference between MGUS and newly diagnosed multiple myeloma. Leukemia 25, 1026–1035, 10.1038/leu.2011.53 (2011). Article
CAS PubMed PubMed Central Google Scholar * Hanamura, I., Huang, Y., Zhan, F., Barlogie, B. & Shaughnessy, J. Prognostic value of cyclin D2 mRNA expression in newly diagnosed
multiple myeloma treated with high-dose chemotherapy and tandem autologous stem cell transplantations. Leukemia 20, 1288–1290, 10.1038/sj.leu.2404253 (2006). Article CAS PubMed Google
Scholar * Pelengaris, S., Khan, M. & Evan, G. c-MYC: more than just a matter of life and death. Nat Rev Cancer 2, 764–776, 10.1038/nrc904 (2002). Article CAS Google Scholar * Korn,
T. et al. IL-21 initiates an alternative pathway to induce proinflammatory T(H)17 cells. Nature 448, 484–487, 10.1038/nature05970 (2007). Article ADS CAS PubMed PubMed Central Google
Scholar * Wang, W. et al. The proteomic investigation reveals interaction of mdig protein with the machinery of DNA double-strand break repair. Oncotarget 6, 28269–28281,
10.18632/oncotarget.4961 (2015). Article PubMed PubMed Central Google Scholar * Cottini, F. et al. Synthetic lethal approaches exploiting DNA damage in aggressive myeloma. Cancer Discov,
10.1158/2159-8290.CD-14-0943 (2015). * Bommert, K., Bargou, R. C. & Stuhmer, T. Signalling and survival pathways in multiple myeloma. Eur J Cancer 42, 1574–1580,
10.1016/j.ejca.2005.12.026 (2006). Article CAS PubMed Google Scholar * Dechow, T. et al. GP130 activation induces myeloma and collaborates with MYC. J Clin Invest 124, 5263–5274,
10.1172/JCI69094 (2014). Article PubMed PubMed Central Google Scholar * Hideshima, T., Mitsiades, C., Tonon, G., Richardson, P. G. & Anderson, K. C. Understanding multiple myeloma
pathogenesis in the bone marrow to identify new therapeutic targets. Nat Rev Cancer 7, 585–598, 10.1038/nrc2189 (2007). Article CAS PubMed Google Scholar * Brown, R. et al. Phospho-flow
detection of constitutive and cytokine-induced pSTAT3/5, pAKT and pERK expression highlights novel prognostic biomarkers for patients with multiple myeloma. Leukemia 29, 483–490,
10.1038/leu.2014.204 (2015). Article CAS PubMed Google Scholar * Ishikawa, H. et al. Accelerated proliferation of myeloma cells by interleukin-6 cooperating with fibroblast growth factor
receptor 3-mediated signals. Oncogene 24, 6328–6332, 10.1038/sj.onc.1208782 (2005). Article CAS PubMed Google Scholar * Loffler, D. et al. Interleukin-6 dependent survival of multiple
myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 110, 1330–1333, 10.1182/blood-2007-03-081133 (2007). Article CAS PubMed
Google Scholar * Xing, J., Wang, K., Liu, P. W., Miao, Q. & Chen, X. Y. Mina53, a novel molecular marker for the diagnosis and prognosis of gastric adenocarcinoma. Oncol Rep 31,
634–640, 10.3892/or.2013.2918 (2014). Article CAS PubMed Google Scholar * Chowdhury, R. et al. Ribosomal oxygenases are structurally conserved from prokaryotes to humans. Nature 510,
422–426, 10.1038/nature13263 (2014). Article ADS CAS PubMed PubMed Central Google Scholar * Kim, E. et al. Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and
promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 23, 839–852, 10.1016/j.ccr.2013.04.008 (2013). Article CAS PubMed PubMed Central Google Scholar * Justilien, V. et
al. The PRKCI and SOX2 oncogenes are coamplified and cooperate to activate Hedgehog signaling in lung squamous cell carcinoma. Cancer Cell 25, 139–151, 10.1016/j.ccr.2014.01.008 (2014).
Article CAS PubMed PubMed Central Google Scholar Download references ACKNOWLEDGEMENTS This work was partially supported by NIH grants R01 ES017217, R01 ES020137 and P30 ES020957 to FC.
YZ is supported by NIH R01DK107666 and R01DK081750. We thank Dr. Kenneth Reuhl at the Environmental and Occupational Health Sciences Institute of the Rutgers University for providing WTC
dust used in this study. AUTHOR INFORMATION AUTHORS AND AFFILIATIONS * Department of Pharmaceutical Sciences, Eugene Applebaum College of Pharmacy and Health Sciences, Wayne State
University, 259 Mack, Avenue, MI 48201, Detroit, USA Kai Wu, Lingzhi Li, Chitra Thakur, Yongju Lu, Xiangmin Zhang, Zhengping Yi & Fei Chen Authors * Kai Wu View author publications You
can also search for this author inPubMed Google Scholar * Lingzhi Li View author publications You can also search for this author inPubMed Google Scholar * Chitra Thakur View author
publications You can also search for this author inPubMed Google Scholar * Yongju Lu View author publications You can also search for this author inPubMed Google Scholar * Xiangmin Zhang
View author publications You can also search for this author inPubMed Google Scholar * Zhengping Yi View author publications You can also search for this author inPubMed Google Scholar * Fei
Chen View author publications You can also search for this author inPubMed Google Scholar CONTRIBUTIONS K.W. performed most of the experiments and made the first draft of the manuscript;
C.T., L.L. and Y.L. performed immunohistochemistry assays; X.Z. and Z.Y. performed proteomics studies; C.F. analyzed the data and made the final draft of the manuscript. ETHICS DECLARATIONS
COMPETING INTERESTS The authors declare no competing financial interests. RIGHTS AND PERMISSIONS This work is licensed under a Creative Commons Attribution 4.0 International License. The
images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included
under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit
http://creativecommons.org/licenses/by/4.0/ Reprints and permissions ABOUT THIS ARTICLE CITE THIS ARTICLE Wu, K., Li, L., Thakur, C. _et al._ Proteomic Characterization of the World Trade
Center dust-activated mdig and c-myc signaling circuit linked to multiple myeloma. _Sci Rep_ 6, 36305 (2016). https://doi.org/10.1038/srep36305 Download citation * Received: 21 July 2016 *
Accepted: 12 October 2016 * Published: 11 November 2016 * DOI: https://doi.org/10.1038/srep36305 SHARE THIS ARTICLE Anyone you share the following link with will be able to read this
content: Get shareable link Sorry, a shareable link is not currently available for this article. Copy to clipboard Provided by the Springer Nature SharedIt content-sharing initiative