A quasi-direct lc-ms/ms-based targeted proteomics approach for mirna quantification via a covalently immobilized dna-peptide probe
- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:
ABSTRACT MicroRNAs (miRNAs) play a vital role in regulating gene expression and are associated with a variety of cancers, including breast cancer. Their distorted and unique expression is a
potential marker in clinical diagnoses and prognoses. Thus, accurate determination of miRNA expression levels is a prerequisite for their applications. However, the assays currently
available for miRNA detection typically require pre-enrichment, amplification and labeling steps, and most of the assays are only semi-quantitative. Therefore, we developed a quasi-direct
liquid chromatography-tandem mass spectrometry (LC-MS/MS)-based targeted proteomics approach to quantify target miRNA by innovatively converting the miRNA signal into the mass response of a
reporter peptide _via_ a covalently immobilized DNA-peptide probe. Specifically, the probe containing the targeted proteomics-selected substrate/reporter peptide, GDRAVQLGVDPFR/AVQLGVDPFR,
and the DNA sequence complementary to the target miRNA (i.e., miR-21) was first immobilized on APMTS modified silica nanoparticles using PDITC. After the immobilized probe was recognized and
hybridized with the target miRNA, the excess probe was degraded using MBN and followed by a trypsin digestion of the hybrids. The reporter peptide was released and quantified using
LC-MS/MS. The obtained LOQ was 5 pM. Finally, the developed assay was used for the quantitative analysis of miR-21 in breast cells and tissue samples. SIMILAR CONTENT BEING VIEWED BY OTHERS
SINGLE-MOLECULE AMPLIFICATION-FREE MULTIPLEXED DETECTION OF CIRCULATING MICRORNA CANCER BIOMARKERS FROM SERUM Article Open access 10 June 2021 ATTOMOLAR SENSITIVITY MICRORNA DETECTION USING
REAL-TIME DIGITAL MICROARRAYS Article Open access 28 September 2022 QUANTIFICATION OF PURIFIED ENDOGENOUS MIRNAS WITH HIGH SENSITIVITY AND SPECIFICITY Article Open access 27 November 2020
INTRODUCTION MicroRNAs (miRNAs) are single-stranded, noncoding RNAs with a typical length of 18–25 nucleotides. They generally take part in a number of biological processes as regulatory
factors _via_ repression of messenger RNA (mRNA) translation1,2,3. Recently, accumulating evidence has demonstrated that miRNA plays a very important role in the development of various
cancers, including breast cancer4,5,6,7. For example, the differential levels of miRNA in healthy volunteers and breast cancer patients indicated the potential use of miRNA as a
diagnostic/prognostic marker and therapeutic target5, 8, 9. Therefore, accurate determination of miRNA expression levels is a prerequisite for their application. The relatively short length
and low abundance of miRNAs are disadvantages that must be overcome in a detection platform10, 11. To date, many detection techniques have been reported for the study of the miRNA expression
profile, including indirect methods (e.g., microarrays12, polymerase chain reaction (PCR)13, 14, next-generation sequencing15, electrocatalysis16 and fluorescent resonance energy transfer
(FRET))17 and direct methods (e.g., differential interference contrast (DIC) imaging18, spectral detection assisted by duplex-specific nuclease10, electrochemical-based methods and capillary
electrophoresis (CE)-based methods)19, 20. Indirect detection methods normally require pre-amplification and chemical or enzymatic modification of the target miRNA. These additional steps
often result in information loss, increased assay time and sequence-related biases during quantification20. Notably, most of these methods made use of fluorescence detection. In a previous
study, the beads array for the detection of gene expression (BADGE) assay immobilized oligonucleotide capture probes on fluorescence microspheres, and the beads were then hybridized to
labeled cRNA and passed through a counting device, which records the identity of each microsphere and its probe intensity21. The subsequent studies mostly combined amplification with
fluorescence detection to increase the sensitivity, e.g., using PCR22 or some enzyme-aided target recycling techniques23. However, fluorescence signals are easily affected by temperature and
pH24. In addition, spectral separation often requires a difference in fluorescence for multiplex analysis, which could be difficult25. Direct detection techniques are promising, but they
are primarily in the early stages of development and still require significant efforts to make them applicable. Furthermore, most assays only provide a limited degree of qualitative data26.
Thus, cross comparison could be difficult among different studies (or different laboratories)27. With the recent increasing demand for the detection of slight deregulations in miRNA levels
rather than the presence or absence of a particular miRNA species, techniques that provide high quantitative accuracy are of great interest. Consequently, a mass spectrometry technique,
liquid chromatography-tandem mass spectrometry (LC-MS/MS)-based targeted proteomics, has attracted our attention. LC-MS/MS-based targeted proteomics has become an indispensable tool for
protein quantification in recent years28,29,30, mainly because of the remarkable performance of modern mass spectrometry (e.g., high sensitivity, high selectivity and wide dynamic range)31,
32. The key concept of this technique is to specifically determine the protein of interest at the peptide level33. Peptides generated from the proteolytic digestion of the target protein
serve as surrogate analytes. Selected or multiple reaction monitoring (SRM or MRM) can be used to quantify the selected surrogate peptides30. However, the application of mass spectrometry to
directly detect miRNA has encountered some difficulties due to the complicated and unresolved mass spectra of miRNA, which was discussed in previous studies34,35,36,37. Within this context,
we introduced the concept of a surrogate peptide into miRNA quantification and developed a targeted proteomics approach38. Specifically, the target miRNA was biotinylated and attached to
streptavidin agarose in advance. Its signal was then converted into a reporter peptide _via_ a self-designed DNA-peptide probe, and the reporter peptide was ultimately quantified using
LC-MS/MS. Similar to other indirect methods, the target miRNA modification is susceptible to several issues. In addition to the previously mentioned issues, the biotinylation kit uses T4 RNA
ligase to conjugate a single nucleotide analog to the 3′ terminus of the RNA strand39. This conjugation reaction is not very efficient (~70%), and both intramolecular and intermolecular
joining of RNA molecules are possible40, 41. Second, even though biotin and streptavidin form one of the strongest non-covalent interactions (with a dissociation constant of ~10−15 M)42, it
is still not a covalent bond. Dissociation is possible in response to a change in the ionic strength and/or temperature43. Third, the ubiquitous presence of biotin in eukaryotic cells could
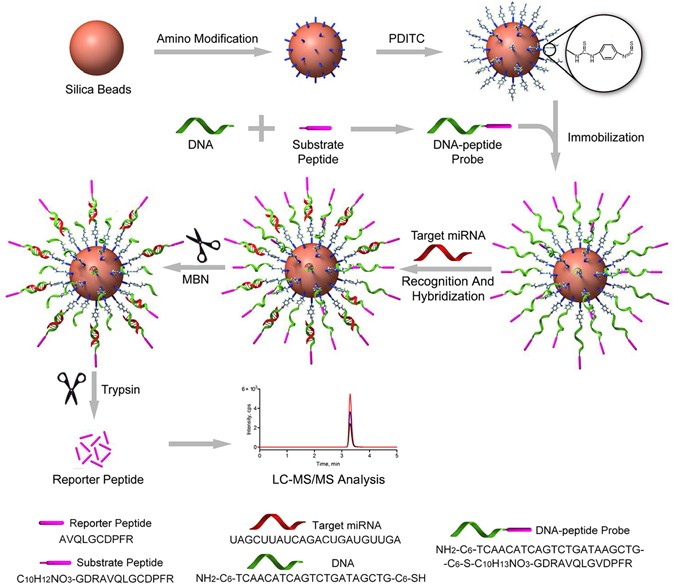
have a significant interfering impact on the miRNA loading efficiency44. In this study, we developed a quasi-direct targeted proteomics approach by covalently immobilizing a DNA-peptide
probe on amino-modified silica nanoparticles (Fig. 1). A substrate peptide containing the reporter peptide and tryptic cleavage site was first designed using a targeted proteomics rationale
and then conjugated with a DNA sequence complementary to the target miRNA. After the _p_-phenylene diisothiocyanate (PDITC) reaction, the newly formed DNA-peptide probe was immobilized on
silica beads that were amino modified using (3-aminopropyl) trimethoxysilane (APTMS) in advance. This immobilized probe was then hybridized with the target miRNA (i.e., miR-21). After
removing the excess probe using mung bean nuclease (MBN), the reporter peptide was liberated from the hybrids using trypsin and ultimately quantified using LC-MS/MS. The new assay for miR-21
was optimized for the parameters including conjugation, immobilization, hybridization and digestion. Finally, miR-21 was quantified in 3 breast cancer cell lines and 36 pairs of human
breast primary tumors and adjacent normal tissue samples. The obtained data were compared with the quantitative reverse transcription PCR (qRT-PCR) results. MATERIALS AND METHODS CHEMICALS
AND REAGENTS Maleimidohexanoic acid-modified substrate peptide (Male-GDRAVQLGVDPF -R), reporter peptide (AVQLGVDPFR) as well as internal standard peptide containing stable-isotope labeled
amino acids were synthesize by ChinaPeptides Co., Ltd. (Shanghai, China). Purification of the peptides was also provided by the manufacturer. The stable isotope-labeled amino acid was
supplied by Cambridge Isotope Laboratories, Inc. (Andover, USA). miR-21 with/without mismatches, and its complementary DNA both with a disulfide modification at 5′ end and amino modification
at 3′ end were custom synthesized by Genscript (Nanjing, China). miR-21 has the following sequence, 5′-UAGCUUAUCAGACUGAUGUUGA-3′. The corresponding DNA is
5′-amino-C6-TCAACATCAGTCTGATAAGCTA-C6-thiol-3′. PDITC was provided by Acros Organics (Geel, Belgium). Silica nanoparticles (40 nm) were supplied by Haotian technology Co., Ltd. (Shanghai,
China). Ammonium bicarbonate (NH4HCO3) was obtained from Qiangshun Chemical Reagent Co., Ltd. (Shanghai, China). Immobilized TCEP reducing beads were obtained from Thermo Scientific
(Rockford, USA). Sequencing grade modified trypsin and MBN with its reaction buffer were both purchased from Promega (Madison, USA). Phosphate buffered saline (PBS) was purchased from
Beyotime Institute of Biotechnology (Nantong, China). Acetonitrile (ACN) and methanol were obtained from Tedia Company, Inc. (Fairfield, USA). Cyanotic chloride, APTMS and trifluoroacetic
acid (TFA) were gained from Aladdin Chemistry Co., Ltd. (Shanghai, China). Diisopropyl ethylamine (DIEA), dimethylformamide (DMF) and 9-fluorenylmethyl chloroformate (Fmoc-Cl) were both
purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China). Formic acid (FA) and sodium borate were obtained from Xilong Chemical Co., Ltd. (Shantou, China). Dulbecco’s Modified
Eagle Media (DMEM), fetal bovine serum and penicillin/streptomycin solutions were obtained from Thermo Scientific HyClone (Logan, USA). Trypan blue and sodium dodecyl sulfate (SDS) were
obtained from Generay Biotech Co., Ltd (Shanghai, China). All the solutions used in the experiments were prepared in DEPC-treated water (Beyotime Institute of Biotechnology, Nantong, China).
PREPARATION OF STOCK SOLUTIONS, CALIBRATION STANDARDS AND QUALITY CONTROLS (QCS) A 100 μM stock solution of miR-21 in DEPC water was prepared and stored in an amber glass tube to protect it
from light at −20 °C. The calibration standards were created by serially diluting the stock solutions in a 10 μM tRNA library from baker’s yeast (Sigma, St. Louis, MO, USA)45. The
concentrations of the calibration standards were 5 pM, 10 pM, 25 pM, 100 pM, 1 nM, and 10 nM. The QC standards for the lower limit of quantification (LLOQ), low QC, mid QC and high QC were
prepared at 5 pM, 15 pM, 500 pM and 8 nM and frozen prior to use. For the reporter peptide, the corresponding isotope-labeled synthetic peptide was used as the internal standard. A 100 μM
internal standard stock solution was prepared, and a 1 nM working solution was prepared by diluting the stock solution with an ACN:water mixture (50:50, _v_/_v_) containing 0.1% FA. CELL
CULTURE AND TISSUE COLLECTION MCF-10A cells (ATCC, Manassas, VA) and MCF-7/WT (ATCC, Manassas, VA) cells were cultured routinely in DMEM media supplemented with 10% fetal bovine serum and 1%
penicillin/streptomycin in a humidified atmosphere with 5% CO2 at 37 °C. MCF-7/ADR (Keygen Biotech, Nanjing, China) cells were grown in an RPMI 1640 media (with L-glutamine and sodium
pyruvate) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin at 37 °C under a 5% CO2 atmosphere. Cells were split every 5–7 days by lifting the cells with 0.25% trypsin,
and the cells were maintained by the addition of fresh medium. The MCF-7/ADR cells were periodically reselected by growing them with 1000 ng/mL DOX to maintain a highly drug-resistant cell
population46. Experiments were conducted using the cells incubated without DOX for 48 h. The cells were counted using a hemocytometer (Qiujing, Shanghai, China). The cell viability was
assessed using trypan blue (0.4%) exclusion. The cell suspensions, PBS and trypan blue were mixed in a 2:3:5 ratio, and the viable cells were counted after incubation for 5 min at 37 °C. The
breast tissue collection in the present study was approved by the ethical review committee of Nanjing Medical University, and informed consent was obtained from all patients. All
experiments were performed in accordance with the relevant guidelines and regulations. Thirty-six pairs of tumors and adjacent normal tissue samples were collected from breast cancer
patients between January 2012 and December 2012 at the First Affiliated Hospital of Nanjing Medical University and Nanjing Drum Tower Hospital, Nanjing, China (mean patient age, 52.9 ± 8.6
years; age range, 38–65 years). The pathologic examinations of the tissue sections were confirmed as normal and/or cancerous by the hospital pathologists. The histological evaluations of the
adjacent tissue samples indicated that there was no contamination from the tumor or other abnormal cells. All the participants were biologically unrelated and belonged to the Han Chinese
ethnic group from the Jiangsu province in China. After collection, the samples were stripped of adhering fat, cut into small pieces, and stored at −80 °C. Before RNA extraction, they were
thawed to room temperature and rinsed thoroughly with DEPC water. The total RNA was isolated from the cells, and using TRIzol Reagent, the concentrations of approximately 50 mg tissue
homogenates were estimated using a nanodrop spectrophotometer (Thermo Fisher Scientific Inc., MA, USA). The RNA extracts were stored at −80 °C until further processing. PREPARATION OF THE
DNA-PEPTIDE PROBE The experimental procedure to prepare the DNA-peptide probe has been described in our previous work38. Briefly, 1 OD DNA (4.55 nmol) was first reduced using 300 μL of TCEP
reducing beads at 37 °C for 2 h with vigorous shaking. After centrifugation at 1000 g for 5 min, the supernatant was collected, and an equal volume of 50 nmol maleimidohexanoic acid-modified
substrate peptide was added to the solution. Prior to purification, the conjugation reaction was performed at 37 °C for 4 h with vigorous shaking. The DNA-peptide compound was separated
from the excess non-conjugated DNA and peptide using high-performance liquid chromatography (HPLC). Finally, the collected fraction was ultra-filtered using an Amicon Ultra 3 K device (Merck
Millipore, Darmstadt, Germany). Quantification was performed using an external calibration peak area measurement. The HPLC conditions are provided in the Supporting Information. AMINO
MODIFICATION OF THE SILICA BEADS AND AMINO LOADING ESTIMATION Silica beads (~50 mg) were washed with ethanol and resuspended in 1 mL of ethanol, followed by an addition of 10 μL (1%) APTMS
and shaking for 1 h at room temperature. After amino modification, the beads were washed 5 times with ethanol and resuspended in 1 mL of methylene chloride, which was followed by an addition
of 20 μL (0.12 mmol) of DIEA and sonication for 5 min. To estimate the available amino groups, 25.8 mg (0.1 mmol) of Fmoc-Cl were added into the suspension, and the mixture was shaken for 1
h at room temperature. Following centrifugation at 5000 rpm for 5 min, the beads were washed with methylene chloride and ACN 5 times, sequentially. In this study, the unreacted amino groups
were capped using a mixture of 0.2 M acetic anhydride and 0.2 M DIEA in methylene chloride (1 mL for 50 mg of beads) with shaking overnight at room temperature. The beads were then washed
with methylene chloride and ACN 5 times again. Finally, the Fmoc protecting group was removed using 1 mL of 20% piperidine in DMF. After shaking for 30 min, the supernatant was collected and
analyzed using HPLC. IMMOBILIZATION OF THE DNA-PEPTIDE PROBE The amino-modified beads (50 mg) were suspended in 1 mL of DMF containing 10% pyridine and 0.2% PDITC and shaken for 2 h at room
temperature. The beads were then washed with DMF 5 times, with ethanol 3 times and with methylene chloride 3 times. Afterward, 0.75 nmol of the DNA-peptide probe and 10 mg of the beads were
mixed in 400 μL of the reaction buffer, which included 2 M sodium chloride and 0.05 M sodium borate buffer at pH 8.5. After shaking overnight at 37 °C, the beads were washed with PBS 3
times. MIRNA HYBRIDIZATION WITH THE DNA-PEPTIDE PROBE After immobilization, 1 nM miR-21 in the hybridization buffer (200 μL) was mixed with 20 μL of the probe immobilized beads (0.25 mg),
and the reaction was conducted in an oven. Similar to previous work38, the hybridization buffer (Table 1S), time and temperature were optimized. After hybridization, the beads were
thoroughly washed to remove any unbound miRNAs. To evaluate the hybridization specificity of the probe, the miR-21 molecules with a single-base mismatch (5′-UAGCUUAUCAGUCUGAUGUUGA-3′) and
two-base mismatch (5′-UAGCUUAUCAGUGUGAUGUUGA-3′) sequences were also examined. The underlined bases in the sequences represent the mismatch sites. IN-SOLUTION MBN DIGESTION AND TRYPTIC
DIGESTION After hybridization, 0.25 mg of the beads were treated with 40 units of MBN for 20 min at room temperature in 200 μL of the reaction buffer containing 30 mM sodium acetate (pH
5.0), 50 mM sodium chloride and 1 mM zinc chloride47, 48. The beads were then washed with 10 mM PBS 3 times. After centrifugation, 200 μL of 50 mM NH4HCO3 were mixed with the beads and
followed by the addition of 1 μg of sequencing grade trypsin and incubation at 37 °C for 24 h. Then, 10 μL of 0.1% TFA were added to stop the reaction. After that, the tryptic mixture, with
an addition of 100 μL of the internal standard solution, was transferred into a microspin C18 column (The Nest Group, Inc., MA, USA). The column was preconditioned using 100 μL of ACN and
100 μL of water in advance. After loading, the column was washed with 50 μL of 5% ACN containing 0.1% TFA and eluted with 50 μL of 80% ACN containing 0.1% FA. The elution was repeated for
another 3 times. For additional detailed information, please see the manufacturer’s operating instruction. LC-MS/MS METHOD DEVELOPMENT AND VALIDATION The LC-MS/MS analysis was performed
using an Agilent Series 1290 UPLC system (Agilent Technologies, Waldbronn, Germany) coupled to a 6460 Triple Quad LC-MS mass spectrometer (Agilent Technologies, Santa Clara, USA).
Chromatographic separation was achieved using an Agilent SB C18 (2.7 µm, 30 mm × 2.1 mm) at room temperature. The analysis was performed in the positive ion mode. The mobile phase A was
0.02% FA in water, and the mobile phase B was 0.02% FA in methanol with a flow rate of 0.3 mL/min. The elution gradient started with 10% of eluent B run at isocratic conditions for 1 min,
and then, it was increased to 90% in 4 min and held for another 4 min. Finally, the initial condition was reached again in 1 min. The sample injection volume was set at 5 μL. The mass
spectrometer was equipped with an electrospray ion source. The following parameters were used: Q1 and Q3: set at unit resolution; the drying gas: 10 L/min, 350 °C; needle voltage: 4000 V;
nebulizer pressure: 35 psi. The data were obtained and processed using the Agilent MassHunter Workstation Software (version B.06.00) The precision and accuracy of the assay were determined
using QC samples. The detailed validation procedures and acceptance criteria for the assay have been discussed in a number of studies31, 49. METHOD COMPARISON The qRT-PCR was conducted for
comparison. For the experimental details, please see the Supporting Information. ETHICAL APPROVAL This study was approved by the Institutional Review Board of Nanjing Medical University,
Nanjing, China. RESULTS AND DISCUSSION A QUASI-DIRECT TARGETED PROTEOMICS APPROACH The goal of this approach was to convert the miRNA signal into a reporter peptide for mass spectrometric
analysis. For the scheme to work, an appropriate reporter/substrate peptide must be found. Then, the N-terminal maleimidohexanoic acid-modified substrate peptide was conjugated with the
3′-end thiol-modified complementary DNA to form the DNA-peptide probe (Fig. 2, reaction 3). Instead of the previous modification of the target miRNA, the DNA-peptide probe was covalently
immobilized on the amino-modified silica nanoparticles _via_ a PDITC reaction in this study (Fig. 2, reactions 1, 2 and 4). After reacting with the target miRNA (i.e., miR-21) under the
optimized conditions, the miRNA-DNA-peptide hybrids were formed. The excess single-stranded DNA-peptide probe was degraded _via_ a single-stranded specific nuclease MBN. Finally, the beads
bearing the hybrids were subject to trypsin digestion. Thus, the reporter peptide could be released from the beads and detected using LC-MS/MS. SELECTION OF THE REPORTER AND SUBSTRATE
PEPTIDES In our previous work, the criteria for the selection of suitable reporter and substrate peptides were extensively described38. While evidence indicated that our previously designed
surrogate/reporter peptide was applicable to targeted proteomics analysis, these peptides may not be suitable for the present work because of the probe immobilization process. To date,
oligonucleotide immobilization has been widely used in a number of important molecular applications50. There are several strategies available for immobilization51. Among them, the most
accessible approach is covalent attachment of oligonucleotides to the solid support52. Normally, oligonucleotides are modified at their end-group and then react with the functional groups on
the solid support53, 54. The combinations of oligonucleotide/surface functional groups that have been reported so far include thiol/acrylamide55, amine/isothiocyanate52, activated
carboxylic acid/amine56, 57, amine/aldehyde58, 59 and epoxide/amine60. Among them, the amine/isothiocyanate reaction under mildly alkaline conditions can provide a high immobilization
efficiency for oligonucleotides, which has been proven by many studies52, 61, 62. Thus, PDITC, a member of the isothiocyanate family, was selected to crosslink the DNA-peptide probe to the
beads. In principle, this cross-linker is capable of reacting with the amino-modified beads at one end and with the DNA-peptide probe at the other end (Fig. 2, reactions 3 and 4)52. Its
preferred reactivity with a primary amine is essential for the product yield63. However, additional consideration is required in this study because of the potential interference from the
peptide component of the probe. As is well known, the primary amino groups of the peptides exist both at the N-terminus (α-amino group) and in the side chain of the lysine residues (ε-amino
group)64. The N-terminal amino group was formerly modified and conjugated with the DNA in the probe formation, whereas the ε-amino group was free and could form a stable phenylthiourea (PTU)
derivative during immobilization, if it exists65. Thus, the previously selected substrate peptide, GDKAVLGVDPFR, was no longer suitable because of the presence of the lysine residue at
position 3. Given the biochemical similarity of lysine and arginine66, 67, the replacement of lysine with arginine was easily accomplished while preserving the original tryptic site. The new
substrate peptide was GDRAVQLGVDPFR, and the corresponding reporter peptide was AVQLGVDPFR. Fortunately, the sequence of the reporter peptide did not match any protein based on a BLAST
search, and the peptide had a high predicted mass response (ESP Predictor (0.90, full score is 1). Experimentally, the maximum response was afforded _via_ its doubly charged ion at _m_/_z_
551.3. Its product ion spectrum is shown in Fig. 3A. The sequence-specific b ions and y ions were observed and were indicative of the precursor peptide. The corresponding stable,
isotope-labeled peptide was also synthesized to serve as an internal standard. For this peptide, the stable, isotope-labeled [D8] Val (V*) was coupled to AVQLGVDPFR at positions 2 and 6 to
form AV*QLGV*DPFR, which yielded a molecular mass shift of 16 Da from the non-labeled peptide and a monoisotopic molecular mass of 1117.6 Da. The retention times for AVQLGVDPFR and its
isotope-labeled peptide were identical (∼3.3 min, Fig. 1S-A). Finally, the MRM transitions afforded by the product ions, b2 _m_/_z_ 171.1, y3 _m_/_z_ 419.2 and y6 _m_/_z_ 690.3, gave the
best signal-to-noise (S/N) and limit of quantification (LOQ) (Fig. 3B). This characteristic pattern was also observed in AV*QLGV*DPFR (Fig. 1S-B). The quantification limit of these MRM
transitions was 5 pM (Fig. 2S). Therefore, the peak areas from the three transitions, _m_/_z_ 551.3 → _m_/_z_ 171.1, _m_/_z_ 551.3 → _m_/_z_ 419.2 and _m_/_z_ 551.3 → _m_/_z_ 690.3, were
summed and used in the following quantitative analysis68. The digestion efficiency of GDRAVQLGVDPFR was calculated by comparing the response ratios of the equimolar peptide before and after
digestion. The estimated value was 99.91% (Fig. 3S). To note, the subsequent conjugated DNA, hybridized miR-21 and the probe-coated silica beads did not have a significant impact on the
digestion (please see the following section). The reason may be that the tryptic site was intentionally designed using the three amino acid residues (GDR) and the C6 spacer (on DNA) away
from the DNA sequence to avoid any potential steric hindrance in the trypsin cleavage. Meanwhile, this design also reduced the potential spatial interference of the peptide tag on the
reactivity of the DNA. PREPARATION AND CHARACTERIZATION OF THE DNA-PEPTIDE PROBE To prepare the DNA-peptide probe, the disulfide at the 3′ end of the DNA was first reduced to a thiol, and,
then, a Michael addition reaction was performed between the reactive thiol and the maleimide at the substrate peptide N-terminus to form the thiol-maleimide linkage (Fig. 2, reaction 3)69.
In addition to the factors previously discussed65, there is another factor that deserves consideration. The newly designed amino-modified 5′ end of the DNA as well as its thiol group at the
3′ end can react with the double bond of the maleimide to form stable carbon-nitrogen and carbon-sulfur bonds, respectively70. Fortunately, there is evidence that the maleimide reaction is
specific for thiols at pH 6.5 ~ 7.5 and for the amino group when the pH is greater than 871. After careful evaluation of the pH effect, 10 mM PBS (pH = 7.35) was selected as the conjugation
buffer in this study. To further rule out the possibility of cross-reactivity (Fig. 4S), we used disulfide-protected DNA to react with the peptide. As shown in Fig. 5S, no new compound was
observed in the HPLC chromatogram. Under the above optimized conditions, almost all the DNA molecules were conjugated with the substrate peptide in the presence of excess peptide. The
product was immediately purified using preparative HPLC. As shown in Fig. 4A, the DNA before/after reduction and the DNA-peptide were detected in the HPLC chromatograms using a detection
wavelength of 260 nm; their retention times were 7.8 min, 16.8 min and 21.4 min, respectively. The peptide was undetectable at this wavelength. To ensure that the DNA-peptide conjugate we
collected did not co-elute with the substrate peptide, a 220 nm detection wavelength was chosen for peptide detection under the same conditions. Figure 4B indicates that the substrate
peptide was not eluted out of the column. For another probe characterization, trypsin was added to the collected fraction. The expected products, DNA-GDR and AVQLGVDPFR (i.e., reporter
peptide), were detected using HPLC and LC-MS/MS, respectively. Consistent with the previous observation38, the retention time of the DNA product (9.25 min in HPLC) was not the same as that
of the premier DNA because of the addition of three amino acid residues from the substrate peptide (Fig. 5A). However, the retention of the peptide product agreed with that of the reporter
peptide (Fig. 5B). Furthermore, the estimated amount of the reporter peptide was not significantly different from the amount of the DNA-peptide reactant, which confirmed that the conjugation
of the DNA to the substrate peptide did not have a significant impact on digestion. OPTIMIZATION OF THE DNA-PEPTIDE PROBE IMMOBILIZATION As mentioned earlier, a variety of immobilization
strategies, including entrapment, adsorption and chemical binding, have been used to attach oligonucleotide to solid supports, such as ceramics, silicon, glass, magnetic beads, nylon and
polymers57, 72,73,74. Among them, covalent immobilization, owning to the involvement of chemical derivatization, is one of the most common approaches. In the present study, the silica beads
were first silanized _via_ APTMS to generate an active amino group prior to the PDITC reaction. OPTIMIZATION OF THE APTMS CONCENTRATION To evaluate the influence of the reactant APTMS
concentration on the availability of amino groups on beads, the reaction was performed with an increase in the APTMS concentration from 0.01% to 2.5%. Because it is normally difficult to
directly detect immobilized amino groups, a derivatizing agent, Fmoc-Cl, that can react with free amino groups was introduced. After attachment, the Fmoc group was removed, and its
absorbance was monitored using a UV detector to estimate the available amino groups69, 75. The results indicated that the amount of the amino groups increased as the concentration of APTMS
increased from 0.1% to 1% and reached a plateau afterward (Fig. 6A). Therefore, 1% APTMS was used in the subsequent experiments. OPTIMIZATION OF THE PROBE CONCENTRATION After determining the
availability of the amino group, the PDITC reaction was performed with various concentrations of the DNA-peptide probe, ranging from 0 μM to 3.75 μM. The immobilization efficiency was
calculated as the input of the DNA-peptide probe divided by the amount of the probe immobilized, which can be estimated from the probe amount left in the solution. As shown in Fig. 6B, there
was a direct and positive relationship between the input and the immobilized probe. However, the level of immobilization efficiency decreased throughout the range. This phenomenon may
result from the steric hindrance of the immobilized probes that block the access of the free probes to the adjacent amino sites52. Taking the immobilization amount and efficiency into
account, we chose 1.875 μM as the probe reaction concentration with 87.3% immobilization efficiency. HYBRIDIZATION EFFICIENCY In this study, the hybridization efficiency was assessed using
the molar ratio of the hybridized miRNA and reactant miRNA76, 77. Its value was influenced by many parameters, including hybridization buffer, temperature and time. Six commonly used buffers
were examined, and a buffer of 10 mM Tris, 100 mM KCl, and 1 mM MgCl2 at pH 7.4 (buffer 3) provided the highest efficiency (Table 1S, Fig. 6S). In addition, the optimum temperature and time
were 62 °C and 12 h, respectively. Under these conditions, the hybridization efficiency reached 90.2%. The distinct nature of miRNA concerning detection specificity is the presence of
highly homologous family members. Their sequences differ by only 1–3 nucleotides10, 11, which may create extra difficulties when it is necessary to distinguish specific miRNAs from their
family members. To evaluate the specificity of this assay, we also applied the assay to single-base mismatched miR-21 (SM) and double-base-mismatched miR-21 (DM). The hybridization
efficiency decreased in the order of matched (100%) > SM (9.6%) > DM (3.5%). As predicted, the non-complementary sample did not display a peak. MUNG BEAN NUCLEASE TREATMENT MBN is a
single-stranded specific nuclease purified from the sprouts of the mung bean, _Vigna radiate_ 72. This enzyme prefers to degrade single-stranded DNA (ssDNA) over double-stranded DNA (dsDNA)
or DNA-RNA complexed by 30,000-fold78, 79. In addition, the enzyme can cleave at a single-nucleotide mismatch when two nucleotide strands are not a perfect match. However, the enzyme can
still potentially degrade double-stranded DNA at very high concentrations78, 80. Thus, the specificity of MBN was evaluated. The results indicated that up to 200 U/mL of MBN can digest ssDNA
completely without affecting dsDNA (Fig. 7S). VALIDATION OF THE QUASI-DIRECT TARGETED PROTEOMICS ASSAY After determining the conditions for the immobilization, hybridization and digestion,
the quasi-direct targeted proteomics assay was validated for further application to biological samples. Calibration curves were generated using a weighted linear regression model with a
weighting factor of _1_/_x_ _2_. The relative peak area ratio between the reporter peptide and the stable isotope-labeled internal standard was plotted against the concentration. For the
assay, the LOQ was 5 pM, and the established concentration range was from 5 pM to 10 nM (Fig. 7). Notably, the LOQ was chosen as the concentration corresponding to the lowest standard of the
calibration curve that gave good accuracy and precision as suggested by FDA81. However, we have to admit that the LOQ is at least 5 pM in neat buffer conditions, but may be higher when
realistic sample matrix is present in the sample. The other results are provided in the Supporting Information (Table 2S). QUANTIFICATION OF MIR-21 IN BREAST CELLS AND TISSUE SAMPLES Using
the assay, the levels of miR-21 were accurately quantified as (7.94 ± 1.62) × 103 copies/cell in MCF-10A cells, (3.06 ± 0.45) × 104 copies/cell in MCF-7/WT and (4.90 ± 0.66) × 104
copies/cell in MCF-7/ADR cells. Their difference was statistically significant (p < 0.05). This result agrees well with the results reported in our previous work38, but all the values are
higher. One of the possible reasons could be the indirect nature of the previous method, i.e., miR-21 was biotinylated and bound to streptavidin agarose. The biotinylation process may cause
information loss of the target miRNA because of the complexity of the biological samples, e.g., the ubiquitous presence of biotin44. Finally, miR-21 was measured using quantitative qRT-PCR
for a comparison (Fig. 8S). The level of miR-21 determined using our assay was slightly higher than that obtained using qRT-PCR, but the difference was not significant. Such uncertainty
could be attributed to the larger variations in the qRT-PCR experiment. qRT-PCR is the most widely reported method and is regarded as a gold standard for quantifying miRNAs because of its
high sensitivity82. However, this method suffers from practical issues such as low efficiency, high false positive rates for amplification, and sophisticated and expensive analysis83.
Currently, most qRT-PCR analyses determine the relative miRNA abundance (often with respect to a non-validated reference miRNA)84. Absolute quantification _via_ qRT-PCR can provide a
quantity of unknowns, but it is labor-intensive85. Minor variations in the reaction components, thermal cycling conditions, and mispriming events during the early stages of the reaction can
lead to large changes in the overall amount of the amplified product86. Furthermore, the subsequent analysis is also mathematically complex83. Thus, our assay is able to provide the miR-21
level in copies per cell while avoiding complicated sample manipulation and data analysis. Moreover, we also performed a quasi-direct targeted proteomics analysis of 36 matched pairs of
breast tissue samples. The amounts of miR-21 were (3.63 ± 1.99) × 108 copies/mg (range: (0.49–8.10) × 108 copies/mg) in normal tissues and (1.23 ± 0.44) × 109 copies/mg (range: (0.23–1.93) ×
109 copies/mg) in tumors (Fig. 8). A two-way comparison using the Mann-Whitney test showed that the tumor samples had a significantly higher level of miR-21 compared to the normal samples
(p < 0.0001). Specifically, approximately a 3.4-fold increase in the concentration of miR-21 was observed in tumors. CONCLUSIONS In this study, we immobilized the DNA-peptide probe on
amino-modified silica beads and developed a quasi-direct targeted proteomics assay for miRNA quantification. Using this assay, the target miR-21 was quantified in 3 breast cell lines and 36
pairs of breast tissue samples. The advantages of the quasi-direct targeted proteomics approach were demonstrated by converting the miRNA signal to the mass response of the reporter peptide.
More importantly, immobilization of the DNA-peptide probe circumvents the miRNA biotinylation and subsequent dependence on the biotin-streptavidin interaction and allows the use of RNA
samples without any further manipulation. Technically, this quasi-direct targeted proteomics method can be easily applied to other miRNAs by replacing the DNA sequence with one that is
complementary to the target miRNA while keeping the reporter peptide the same. However, the parameters, including conjugation, immobilization, hybridization and digestion, deserve careful
optimization to achieve the highest sensitivity and specificity for each miRNA. Furthermore, this assay has more potential for simultaneous detection of multiple miRNAs. Indeed, a key
advantage of the LC-MS/MS-based quasi-targeted proteomics assay is its multiplexing ability, which is valid as long as a mass spectrometer can manage the concomitant analysis of multiple
reporter peptides while retaining a degree of selectivity. However, the challenges of optimizing the assay format for each peptide, selecting a common dilution factor, addressing the
variability and cross-interference, and establishing a robust quality control algorithm are substantial and require further analytical and statistical development. We anticipate that the
quasi-targeted proteomics approach described here can ultimately be applied to the profiling of miRNAs in biological samples. This type of quasi-direct analysis may increase the quantitative
accuracy and precision, but more evidence is required to confirm this feature. REFERENCES * Zeng, X., Zhang, X. & Zou, Q. Integrative approaches for predicting microRNA function and
prioritizing disease-related microRNA using biological interaction networks. _Brief. Bioinform._ 17, 193–203 (2016). Article PubMed Google Scholar * Etheridge, A., Lee, I., Hood, L.,
Galas, D. & Wang, K. Extracellular microRNA: a new source of biomarkers. _Mutat. Res._ 717, 85–90 (2011). Article CAS PubMed PubMed Central Google Scholar * Zhao, Y. &
Srivastava, D. A developmental view of microRNA function. _Trends Biochem. Sci._ 32, 189–97 (2007). Article CAS PubMed Google Scholar * Konishi, H. _et al_. Detection of gastric
cancer-associated microRNAs on microRNA microarray comparing pre- and post-operative plasma. _Br J. Cancer_ 106, 740–7 (2012). Article CAS PubMed PubMed Central Google Scholar * Yan, L.
X. _et al_. MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. _RNA_ 14, 2348–60 (2008).
Article CAS PubMed PubMed Central Google Scholar * Tavazoie, S. F. _et al_. Endogenous human microRNAs that suppress breast cancer metastasis. _Nature_ 451, 147–52 (2008). Article CAS
PubMed PubMed Central ADS Google Scholar * Lu, J. _et al_. MicroRNA expression profiles classify human cancers. _Nature_ 435, 834–8 (2005). Article CAS PubMed ADS Google Scholar *
Fix, L. N., Shah, M., Efferth, T., Farwell, M. A. & Zhang, B. MicroRNA expression profile of MCF-7 human breast cancer cells and the effect of green tea polyphenon-60. _Cancer Genom.
Proteom_ 7, 261–77 (2010). CAS Google Scholar * Usmani, A., Shoro, A. A., Memon, Z., Hussain, M. & Rehman, R. Diagnostic, prognostic and predictive value of MicroRNA-21 in breast
cancer patients, their daughters and healthy individuals. _Am. J. Cancer Res_ 5, 2484–90 (2015). PubMed PubMed Central Google Scholar * Yin, B. C., Liu, Y. Q. & Ye, B. C. One-step,
multiplexed fluorescence detection of microRNAs based on duplex-specific nuclease signal amplification. _J. Am. Chem. Soc._ 134, 5064–7 (2012). Article CAS PubMed Google Scholar * Duan,
R. _et al_. Lab in a tube: ultrasensitive detection of microRNAs at the single-cell level and in breast cancer patients using quadratic isothermal amplification. _J. Am. Chem. Soc._ 135,
4604–7 (2013). Article CAS PubMed Google Scholar * Lu, J. & Tsourkas, A. Imaging individual microRNAs in single mammalian cells _in situ_. _Nucleic Acids Res_ 37, e100 (2009).
Article PubMed PubMed Central Google Scholar * Zhang, P. _et al_. Highly sensitive and specific multiplexed microRNA quantification using size-coded ligation chain reaction. _Anal.
Chem._ 86, 1076–82 (2014). Article CAS PubMed Google Scholar * Zhang, P. _et al_. Multiplex ligation-dependent probe amplification (MLPA) for ultrasensitive multiplexed microRNA
detection using ribonucleotide-modified DNA probes. _Chem. Commun (Camb)_ 49, 10013–5 (2013). Article CAS ADS Google Scholar * Wyman, S. K. _et al_. Repertoire of microRNAs in epithelial
ovarian cancer as determined by next generation sequencing of small RNA cDNA libraries. _PLoS One_ 4, e5311 (2009). Article PubMed PubMed Central ADS Google Scholar * Gao, Z. &
Yang, Z. Detection of microRNAs using electrocatalytic nanoparticle tags. _Anal. Chem._ 78, 1470–7 (2006). Article CAS PubMed Google Scholar * Jin, Z., Geißler, D., Qiu, X., Wegner, K.
D. & Hildebrandt, N. A Rapid, Amplification-Free, and Sensitive Diagnostic Assay for Single-Step Multiplexed Fluorescence Detection of MicroRNA. _Angew. Chem. Int. Edit_ 54, 10024–9
(2015). Article CAS Google Scholar * Roy, S., Soh, J. H. & Gao, Z. A microfluidic-assisted microarray for ultrasensitive detection of miRNA under an optical microscope. _Lab Chip_ 11,
1886–94 (2011). Article CAS PubMed Google Scholar * Wegman, D. W., Cherney, L. T., Yousef, G. M. & Krylov, S. N. Universal drag tag for direct quantitative analysis of multiple
microRNAs. _Anal. Chem._ 85, 6518–23 (2013). Article CAS PubMed Google Scholar * Wegman, D. W. & Krylov, S. N. Direct miRNA-hybridization assays and their potential in diagnostics.
_TrAC Trend. Anal. Chem._ 44, 121–30 (2013). Article CAS Google Scholar * Yang, L., Tran, D. K. & Wang, X. BADGE, BeadsArray for the detection of gene expression, a high-throughput
diagnostic bioassay. _Genome res._ 11, 1888–98 (2001). CAS PubMed PubMed Central Google Scholar * Peck, D. _et al_. A method for high-throughput gene expression signature analysis.
_Genome bio._ 7, R61 (2006). Article Google Scholar * Lu, J., Paulsen, I. T. & Jin, D. Application of exonuclease III-aided target recycling in flow cytometry: DNA detection
sensitivity enhanced by orders of magnitude. _Anal. Chem._ 85, 8240–5 (2013). Article CAS PubMed Google Scholar * Pospı́šil, P., Skotnica, J. & Nauš, J. Low and high temperature
dependence of minimum F 0 and maximum F M chlorophyll fluorescence _in vivo_. _Biochimica et Biophysica Acta (BBA)-Bioenergetics_ 1363, 95–9 (1998). Article Google Scholar * Riley, J. A.,
Brown, T., Gale, N., Herniman, J. & Langley, G. J. Self-reporting hybridisation assay for miRNA analysis. _Analyst_ 139, 1088–92 (2014). Article CAS PubMed ADS Google Scholar *
Marx, H. _et al_. A large synthetic peptide and phosphopeptide reference library for mass spectrometry-based proteomics. _Nat. Biotechnol._ 31, 557–64 (2013). Article CAS PubMed Google
Scholar * Degliangeli, F., Kshirsagar, P., Brunetti, V., Pompa, P. P. & Fiammengo, R. Absolute and direct microRNA quantification using DNA-gold nanoparticle probes. _J. Am. Chem. Soc._
136, 2264–7 (2014). Article CAS PubMed Google Scholar * Doerr, A. Targeted proteomics. _Nat. Methods._ 7, 34 (2010). Article CAS Google Scholar * Balogh, L. M., Kimoto, E., Chupka,
J., Zhang, H. & Lai, Y. Membrane Protein Quantification by Peptide-Based Mass Spectrometry Approaches: Studies on the Organic Anion-Transporting Polypeptide Family. _J_. _Proteomics
Bioinform_. S4 (2012). * Calvo, E., Camafeita, E., Fernández-Gutiérrez, B. & López, J. A. Applying selected reaction monitoring to targeted proteomics. _Expert Rev. Proteomics._ 8,
165–73 (2011). Article CAS PubMed Google Scholar * Thomas, B. & Akoulitchev, A. V. Mass spectrometry of RNA. _Trends Biochem. Sci._ 31, 173–81 (2006). Article CAS PubMed Google
Scholar * Kiyonami, R. _et al_. Increased selectivity, analytical precision, and throughput in targeted proteomics. _Mol_. _Cell Proteomics_. 10, M110.002931 (2011). * Elschenbroich, S.
& Kislinger, T. Targeted proteomics by selected reaction monitoring mass spectrometry: applications to systems biology and biomarker discovery. _Mol. Biosyst._ 7, 292–303 (2011). Article
CAS PubMed Google Scholar * Thompson, A. _et al_. Electrospray ionisation-cleavable tandem nucleic acid mass tag-peptide nucleic acid conjugates: synthesis and applications to
quantitative genomic analysis using electrospray ionisation-MS/MS. _Nucleic Acids Res._ 35, e28 (2007). Article PubMed PubMed Central Google Scholar * Schürch, S., Bernal-Méndez, E.
& Leumann, C. J. Electrospray tandem mass spectrometry of mixed-sequence RNA/DNA oligonucleotides. _J. Am. Soc. Mass Spectrom._ 13, 936–45 (2002). Article PubMed Google Scholar * Ono,
T., Scalf, M. & Smith, L. M. 2′-Fluoro modified nucleic acids: polymerase-directed synthesis, properties and stability to analysis by matrix-assisted laser desorption/ionization mass
spectrometry. _Nucleic Acids Res._ 25, 4581–8 (1997). Article CAS PubMed PubMed Central Google Scholar * Tang, K. _et al_. Matrix-assisted laser desorption/ionization mass spectrometry
of immobilized duplex DNA probes. _Nucleic Acids Res._ 23, 3126–31 (1995). Article CAS PubMed PubMed Central Google Scholar * Xu, F., Yang, T. & Chen, Y. Quantification of microRNA
by DNA-Peptide Probe and Liquid Chromatography-Tandem Mass Spectrometry-Based Quasi-Targeted Proteomics. _Anal. Chem._ 88, 754–63 (2016). Article CAS PubMed Google Scholar *
http://www.thermofisher.com/order/catalog/product/20160 (Date of access: 28th Mar 2017). * Moore, M. J. & Sharp, P. A. Site-specific modification of pre-mRNA: the 2′-hydroxyl groups at
the splice sites. _Science_ 256, 992–7 (1992). Article CAS PubMed ADS Google Scholar * Romaniuk, PaU. _Methods in Enzymology_. In R. Wu, L. Grossman and K. Moldave(Ed.). 100, 52–56
(1983). * Wilchek, M. & Bayer, E. A. The avidin-biotin complex in bioanalytical applications. _Anal. Biochem._ 171, 1–32 (1988). Article CAS PubMed Google Scholar * Egli, S.,
Schlaad, H., Bruns, N. & Meier, W. Functionalization of block copolymer vesicle surfaces. _Polymers (Basel)._ 3, 252–280 (2011). Google Scholar * İLÇÖL, İLÇÖL & Taga, Y. The Use of
a Nonradioactive Digoxigenin Labeled Probe for Detection of Apo (a) Gene Size. _Turk_. _J_. _Med_. _Sci_. 30, 247-252 (2000). * Redshaw, N. _et al_. Materials and methods RNA samples.
_Biotechniques_ 54, 155–64 (2013). Article CAS PubMed Google Scholar * van den Brand, M. _et al_. Sequential immunohistochemistry: a promising new tool for the pathology laboratory.
_Histopathology._ 65, 651–7 (2014). Article PubMed Google Scholar * Hou, S. Y., Hsiao, Y. L., Lin, M. S., Yen, C. C. & Chang, C. S. MicroRNA detection using lateral flow nucleic acid
strips with gold nanoparticles. _Talanta_ 99, 375–9 (2012). Article CAS PubMed Google Scholar * Murray, M. G. Use of sodium trichloroacetate and mung bean nuclease to increase
sensitivity and precision during transcript mapping. _Anal. Biochem._ 158, 165–70 (1986). Article CAS PubMed Google Scholar * Barnidge, D. R. _et al_. Absolute quantification of the G
protein-coupled receptor rhodopsin by LC/MS/MS using proteolysis product peptides and synthetic peptide standards. _Anal. Chem._ 75, 445–51 (2003). Article CAS PubMed Google Scholar *
Zammatteo, N. _et al_. Comparison between different strategies of covalent attachment of DNA to glass surfaces to build DNA microarrays. _Anal. Biochem._ 280, 143–50 (2000). Article CAS
PubMed Google Scholar * Rao, K. S. _et al_. A novel route for immobilization of oligonucleotides onto modified silica nanoparticles. _Anal. Chim. Acta_ 576, 177–83 (2006). Article CAS
PubMed Google Scholar * Steinberg, G., Stromsborg, K., Thomas, L., Barker, D. & Zhao, C. Strategies for covalent attachment of DNA to beads. _Biopolymers_ 73, 597–605 (2004). Article
CAS PubMed Google Scholar * Nonglaton, G. _et al_. New approach to oligonucleotide microarrays using zirconium phosphonate-modified surfaces. _J. Am. Chem. Soc._ 126, 1497–502 (2004).
Article CAS PubMed Google Scholar * Dugas, V. _et al_. Immobilization of single-stranded DNA fragments to solid surfaces and their repeatable specific hybridization: covalent binding or
adsorption. _Sensor. Actuat. B-Chem_ 101, 112–21 (2004). Article CAS Google Scholar * Chrisey, L. A., Lee, G. U. & O’Ferrall, C. E. Covalent attachment of synthetic DNA to
self-assembled monolayer films. _Nucleic Acids Res_ 24, 3031–9 (1996). Article CAS PubMed PubMed Central Google Scholar * Demers, L. M., Park, S. J., Taton, T. A., Li, Z. & Mirkin,
C. A. Orthogonal assembly of nanoparticle building blocks on dip-pen nanolithographically generated templates of DNA. _Angew. Chem. Int. Edit._ 40, 3071–3 (2001). Article CAS Google
Scholar * Beier, M. & Hoheisel, J. D. Versatile derivatisation of solid support media for covalent bonding on DNA-microchips. _Nucleic Acids Res._ 27, 1970–7 (1999). Article CAS
PubMed PubMed Central Google Scholar * MacBeath, G. & Schreiber, S. L. Printing proteins as microarrays for high-throughput function determination. _Science_ 289, 1760–3 (2000). CAS
PubMed ADS Google Scholar * Guschin, D. _et al_. Manual manufacturing of oligonucleotide, DNA, and protein microchips. _Anal. Biochem._ 250, 203–11 (1997). Article CAS PubMed Google
Scholar * Osborne, M. A. _et al_. Probing DNA Surface Attachment and Local Environment Using Single Molecule Spectroscopy. _J. Phys. Chem., B_ 105, 3120–3126 (2001). Article CAS Google
Scholar * Manning, M. _et al_. A versatile multi-platform biochip surface attachment chemistry. _Mater. Sci. Eng., C_ 23, 347–351 (2003). Article Google Scholar * Manning, M. &
Redmond, G. Formation and characterization of DNA microarrays at silicon nitride substrates. _Langmuir._ 21, 395–402 (2005). Article CAS PubMed Google Scholar * Bąchor, R., Kluczyk, A.,
Stefanowicz, P. & Szewczuk, Z. New method of peptide cleavage based on Edman degradation. _Mol Divers._ 17, 605–11 (2013). Article PubMed PubMed Central Google Scholar * Freed, J.
K., Smith, J. R., Li, P. & Greene, A. S. Isolation of signal transduction complexes using biotin and crosslinking methodologies. _Proteomics._ 7, 2371–4 (2007). Article CAS PubMed
Google Scholar * Edman, P. Identification and semiquantitative determination of phenyl thiohydantoins. _Acta Chem. Scand._ 10, 1507–1509 (1956). Article CAS Google Scholar * Simpson,
R.J. Fragmentation of protein using trypsin. _CSH Protoc_. 2006, pdb. prot4550 (2006). * Olsen, J. V., Ong, S. E. & Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine
residues. _Mol. Cell. Proteomics_ 3, 608–14 (2004). Article CAS PubMed Google Scholar * Kuhn, E. _et al_. Quantification of C-reactive protein in the serum of patients with rheumatoid
arthritis using multiple reaction monitoring mass spectrometry and 13C-labeled peptide standards. _Proteomics_ 4, 1175–86 (2004). Article CAS PubMed Google Scholar * Nitin, N.,
Santangelo, P. J., Kim, G., Nie, S. & Bao, G. Peptide-linked molecular beacons for efficient delivery and rapid mRNA detection in living cells. _Nucleic Acids Res_ 32, e58 (2004).
Article PubMed PubMed Central Google Scholar * Sharpless, N. E. & Flavin, M. The reactions of amines and amino acids with maleimides. Structure of the reaction products deduced from
infrared and nuclear magnetic resonance spectroscopy. _Biochemistry._ 5, 2963–71 (1966). Article CAS PubMed Google Scholar * Brewer, C. F. & Riehm, J. P. Evidence for possible
nonspecific reactions between N-ethylmaleimide and proteins. _Anal. Biochem._ 18, 248–255 (1967). Article CAS Google Scholar * Zammatteo, N. _et al_. Comparison between microwell and bead
supports for the detection of human cytomegalovirus amplicons by sandwich hybridization. _Anal. Biochem._ 253, 180–9 (1997). Article CAS PubMed Google Scholar * Lee, P. H., Sawan, S.
P., Modrusan, Z., Arnold, L. J. & Reynolds, M. A. An efficient binding chemistry for glass polynucleotide microarrays. _Bioconjug. Chem._ 13, 97–103 (2002). Article CAS PubMed Google
Scholar * Lindroos, K., Liljedahl, U., Raitio, M. & Syvänen, A. C. Minisequencing on oligonucleotide microarrays: comparison of immobilisation chemistries. _Nucleic Acids Res._ 29,
E69–9 (2001). Article CAS PubMed PubMed Central Google Scholar * Lubell, W. D. Peptide chemistry. _J. Org. Chem._ 77, 7137–42 (2012). Article CAS PubMed Google Scholar * Gingeras,
T. R., Kwoh, D. Y. & Davis, G. R. Hybridization properties of immobilized nucleic acids. _Nucleic Acids Res._ 15, 5373–90 (1987). Article CAS PubMed PubMed Central Google Scholar *
Liu, Z. C., Zhang, X., He, N. Y., Lu, Z. H. & Chen, Z. C. Probing DNA hybridization efficiency and single base mismatch by X-ray photoelectron spectroscopy. _Colloids Surf. B.,
Biointerfaces._ 71, 238–42 (2009). Article CAS PubMed Google Scholar * Kroeker, W. D., Kowalski, D. & Laskowski, M. Mung bean nuclease I. Terminally directed hydrolysis of native
DNA. _Biochemistry_ 15, 4463–7 (1976). Article CAS PubMed Google Scholar * Desai, N. A. & Shankar, V. Single-strand-specific nucleases. _FEMS Microbiol. Rev._ 26, 457–91 (2003).
Article CAS PubMed Google Scholar * Ardelt, W. & Laskowski, M. Mung bean nuclease I. IV. An improved method of preparation. _Biochem. Biophys. Res. Commun._ 44, 1205–11 (1971).
Article CAS PubMed Google Scholar * Biopharmaceutics Coordinating Committee in CDER, Guidance for industry-bioanalytical method validation, available at:
http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/ucm070107.pdf (Date of access: 28th Mar 2017) (2001). * Mauroy, A., V der Poel, W. H., der Honing, R. H.,
Thys, C. & Thiry, E. Development and application of a SYBR green RT-PCR for first line screening and quantification of porcine sapovirus infection. _BMC Vet. Res._ 8, 193–204 (2012).
Article CAS PubMed PubMed Central Google Scholar * Wong, M. L. & Medrano, J. F. Real-time PCR for mRNA quantitation. _Biotechniques_ 39, 75–85 (2005). Article CAS PubMed Google
Scholar * Wang, T., Viennois, E., Merlin, D. & Wang, G. Microelectrode miRNA sensors enabled by enzymeless electrochemical signal amplification. _Anal. Chem._ 87, 8173–80 (2015).
Article CAS PubMed Google Scholar * Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and
sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. _Biotechnol. Lett._ 26, 509–15 (2004). Article CAS PubMed Google Scholar * Wu, D. Y., Ugozzoli, L., Pal, B.
K., Qian, J. & Wallace, R. B. The effect of temperature and oligonucleotide primer length on the specificity and efficiency of amplification by the polymerase chain reaction. _DNA Cell
Biol._ 10, 233–8 (1991). Article CAS PubMed Google Scholar Download references ACKNOWLEDGEMENTS The National Natural Science Fund (21675089, 21175071), the Project sponsored by SRF for
ROCS, SEM (39), the Jiangsu Six-type Top Talents Program (D) and the Open Foundation of Nanjing University (SKLACLS1102) awarded to Dr. Chen and the Postgraduate Research & Practice
Innovation Program of Jiangsu Province for Mr. Liu are gratefully acknowledged. AUTHOR INFORMATION AUTHORS AND AFFILIATIONS * School of Pharmacy, Nanjing Medical University, Nanjing, 211166,
China Liang Liu, Qingqing Xu, Shuai Hao & Yun Chen Authors * Liang Liu View author publications You can also search for this author inPubMed Google Scholar * Qingqing Xu View author
publications You can also search for this author inPubMed Google Scholar * Shuai Hao View author publications You can also search for this author inPubMed Google Scholar * Yun Chen View
author publications You can also search for this author inPubMed Google Scholar CONTRIBUTIONS L.L. and Y.C. participated in the research design. L.L., Q.X. and H.S. conducted the
experiments. Y.C. drafted the manuscript. All authors read and approved the final manuscript. CORRESPONDING AUTHOR Correspondence to Yun Chen. ETHICS DECLARATIONS COMPETING INTERESTS The
authors declare that they have no competing interests. ADDITIONAL INFORMATION PUBLISHER'S NOTE: Springer Nature remains neutral with regard to jurisdictional claims in published maps
and institutional affiliations. ELECTRONIC SUPPLEMENTARY MATERIAL SUPPLEMENTARY INFORMATION RIGHTS AND PERMISSIONS OPEN ACCESS This article is licensed under a Creative Commons Attribution
4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and
the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s
Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not
permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit
http://creativecommons.org/licenses/by/4.0/. Reprints and permissions ABOUT THIS ARTICLE CITE THIS ARTICLE Liu, L., Xu, Q., Hao, S. _et al._ A Quasi-direct LC-MS/MS-based Targeted Proteomics
Approach for miRNA Quantification _via_ a Covalently Immobilized DNA-peptide Probe. _Sci Rep_ 7, 5669 (2017). https://doi.org/10.1038/s41598-017-05495-7 Download citation * Received: 30
January 2017 * Accepted: 30 May 2017 * Published: 18 July 2017 * DOI: https://doi.org/10.1038/s41598-017-05495-7 SHARE THIS ARTICLE Anyone you share the following link with will be able to
read this content: Get shareable link Sorry, a shareable link is not currently available for this article. Copy to clipboard Provided by the Springer Nature SharedIt content-sharing
initiative